 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_016494505.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Nicotianoideae; Nicotianeae; Nicotiana
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 335aa MW: 38123.8 Da PI: 7.0946 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 174.2 | 3.7e-54 | 12 | 141 | 1 | 128 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLpk..kvka.eekewyfFskrdkkyatgkrknratksgyWkatgkdkev 92
+ppGfrFhPtdeel+ +yL+kkv+ + ++l +vi+evd++k+ePwdL++ ++ + ++ewyfFs++dkky+tg+r+nrat++g+Wkatg+dk++
XP_016494505.1 12 VPPGFRFHPTDEELLYYYLRKKVSYEAIDL-DVIREVDLNKLEPWDLKEicRIGSgPQNEWYFFSHKDKKYPTGTRTNRATTAGFWKATGRDKAI 105
69****************************.9***************954555442556************************************ PP
NAM 93 lskkgelvglkktLvfykgrapkgektdWvmheyrl 128
+ ++++ +g++ktLvfy+grap+g+ktdW+mheyrl
XP_016494505.1 106 YLSNSKRIGMRKTLVFYTGRAPHGQKTDWIMHEYRL 141
**********************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 9.81E-61 | 4 | 164 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 58.342 | 12 | 164 | IPR003441 | NAC domain |
| Pfam | PF02365 | 4.4E-28 | 13 | 141 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003002 | Biological Process | regionalization | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009834 | Biological Process | plant-type secondary cell wall biogenesis | ||||
| GO:0010455 | Biological Process | positive regulation of cell fate commitment | ||||
| GO:0048829 | Biological Process | root cap development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 335 aa Download sequence Send to blast |
MMSGAGNGQL SVPPGFRFHP TDEELLYYYL RKKVSYEAID LDVIREVDLN KLEPWDLKEI 60 CRIGSGPQNE WYFFSHKDKK YPTGTRTNRA TTAGFWKATG RDKAIYLSNS KRIGMRKTLV 120 FYTGRAPHGQ KTDWIMHEYR LDDNTAEVEP LQEDGWVVCR VFKKKNYTRG FQHELGEDDQ 180 EQQLHFVDHM KSGSSDPKQN MQQPHQQACY DYASNFDGSM HLPQLLSPDL LSVNPCPQLP 240 LNANMNLNEC SHQNLLRLTS AAGAAGCSSS TFNIIPQHDK FSGDWSFLDK LLASHQGALQ 300 SQAAIGDHLV NPTGTQKYPV FHHHGLDPDL FKFSK |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ut4_A | 2e-50 | 9 | 166 | 14 | 167 | NO APICAL MERISTEM PROTEIN |
| 1ut4_B | 2e-50 | 9 | 166 | 14 | 167 | NO APICAL MERISTEM PROTEIN |
| 1ut7_A | 2e-50 | 9 | 166 | 14 | 167 | NO APICAL MERISTEM PROTEIN |
| 1ut7_B | 2e-50 | 9 | 166 | 14 | 167 | NO APICAL MERISTEM PROTEIN |
| 4dul_A | 2e-50 | 9 | 166 | 14 | 167 | NAC domain-containing protein 19 |
| 4dul_B | 2e-50 | 9 | 166 | 14 | 167 | NAC domain-containing protein 19 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription regulator. Together with BRN1 and BRN2, regulates cellular maturation of root cap. Represses stem cell-like divisions in the root cap daughter cells, and thus promotes daughter cell fate. Inhibits expression of its positive regulator FEZ in a feedback loop for controlled switches in cell division plane. Promotes the expression of genes involved in secondary cell walls (SCW) biosynthesis. {ECO:0000269|PubMed:19081078, ECO:0000269|PubMed:20197506}. | |||||
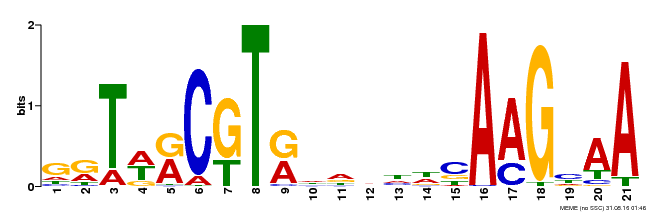
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00250 | DAP | Transfer from AT1G79580 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By FEZ in oriented-divised root cap stem cells. {ECO:0000269|PubMed:19081078}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | HG975524 | 5e-82 | HG975524.1 Solanum lycopersicum chromosome ch12, complete genome. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009794036.1 | 0.0 | PREDICTED: NAC domain-containing protein 76 isoform X1 | ||||
| Refseq | XP_009794037.1 | 0.0 | PREDICTED: NAC domain-containing protein 76 isoform X1 | ||||
| Refseq | XP_016494505.1 | 0.0 | PREDICTED: NAC domain-containing protein 76 isoform X1 | ||||
| Refseq | XP_016494506.1 | 0.0 | PREDICTED: NAC domain-containing protein 76 isoform X1 | ||||
| Swissprot | Q9MA17 | 1e-112 | SMB_ARATH; Protein SOMBRERO | ||||
| TrEMBL | A0A1S4C029 | 0.0 | A0A1S4C029_TOBAC; NAC domain-containing protein 76 isoform X1 | ||||
| TrEMBL | A0A1U7XWP5 | 0.0 | A0A1U7XWP5_NICSY; NAC domain-containing protein 76 isoform X1 | ||||
| STRING | XP_009794036.1 | 0.0 | (Nicotiana sylvestris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA134 | 24 | 297 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G79580.3 | 1e-103 | NAC family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



