 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_016491072.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Nicotianoideae; Nicotianeae; Nicotiana
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 274aa MW: 31288.8 Da PI: 6.0468 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 23 | 1.8e-07 | 21 | 65 | 4 | 46 |
S-HHHHHHHHHHHHHTTTT...-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 4 WTteEdellvdavkqlGgg...tWktIartmgkgRtlkqcksrwqk 46
WT+eE +l+ a++ + + +W +a +++ g++ ++ +q
XP_016491072.1 21 WTKEENKLFESAIAIYDENtpnRWFQVADMIP-GKSVLDVMKQYQE 65
******************9*************.*******999996 PP
| |||||||
| 2 | Myb_DNA-binding | 44.2 | 4.4e-14 | 117 | 161 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+WT+eE+ +++ + ++G+g+W+ I+++m Rt+ q+ s+ qky
XP_016491072.1 117 PWTEEEHRRFLMGLDKYGKGDWRNISKKMVISRTPTQVASHAQKY 161
8*****************************99************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51293 | 7.345 | 16 | 71 | IPR017884 | SANT domain |
| SMART | SM00717 | 3.8E-9 | 17 | 69 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 3.11E-11 | 19 | 73 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 6.1E-6 | 21 | 65 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.12E-7 | 21 | 66 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 1.3E-12 | 108 | 166 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 21.106 | 110 | 166 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.35E-16 | 111 | 165 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 5.4E-18 | 113 | 165 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 4.7E-12 | 114 | 164 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.0E-12 | 117 | 161 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 7.54E-10 | 117 | 161 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 274 aa Download sequence Send to blast |
METIFPISSS WMMQQNKSSI WTKEENKLFE SAIAIYDENT PNRWFQVADM IPGKSVLDVM 60 KQYQELAADV CDIEAGLVPN PGYFASSVTL ELVDDCGLQT FRKRGRSSDQ ERKKGVPWTE 120 EEHRRFLMGL DKYGKGDWRN ISKKMVISRT PTQVASHAQK YYQRQLSGCK DKRRPSIHDI 180 TTFHFVADTS GNNINSLSKE KFYSTPLQKF TTTTDVVNYW NSSSDEDLMD FGSSYGSSVV 240 AYPSEITSQS WDGHGANGNS QLQIQSTRYE IWDS |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 101 | 113 | RKRGRSSDQERKK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Involved in the dorsovental asymmetry of flowers. Promotes ventral identity. {ECO:0000269|PubMed:11937495, ECO:0000269|PubMed:9118809}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00347 | DAP | Transfer from AT3G11280 | Download |
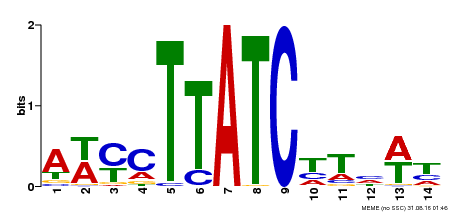
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009627320.1 | 0.0 | PREDICTED: transcription factor DIVARICATA-like | ||||
| Refseq | XP_016491072.1 | 0.0 | PREDICTED: transcription factor DIVARICATA-like | ||||
| Swissprot | Q8S9H7 | 2e-78 | DIV_ANTMA; Transcription factor DIVARICATA | ||||
| TrEMBL | A0A1S4BQA2 | 0.0 | A0A1S4BQA2_TOBAC; transcription factor DIVARICATA-like | ||||
| STRING | XP_009627320.1 | 0.0 | (Nicotiana tomentosiformis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA458 | 24 | 140 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G11280.2 | 2e-82 | MYB family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



