 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_016464627.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Nicotianoideae; Nicotianeae; Nicotiana
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 387aa MW: 42536.2 Da PI: 5.3833 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 49.6 | 7.3e-16 | 141 | 187 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
hn E+rRR+riN+++ L+ l+P++ K +Ka++L +A+eY+k+Lq
XP_016464627.1 141 VHNLSEKRRRSRINEKMKALQNLIPNS-----NKTDKASMLDEAIEYLKQLQ 187
6*************************7.....5******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF47459 | 2.22E-20 | 134 | 197 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 6.2E-20 | 134 | 197 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 17.86 | 137 | 186 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 4.45E-17 | 140 | 191 | No hit | No description |
| Pfam | PF00010 | 2.5E-13 | 141 | 187 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 8.3E-18 | 143 | 192 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010114 | Biological Process | response to red light | ||||
| GO:0010154 | Biological Process | fruit development | ||||
| GO:0010187 | Biological Process | negative regulation of seed germination | ||||
| GO:0048440 | Biological Process | carpel development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 387 aa Download sequence Send to blast |
MANNNIYHHN ASSSFSLTEP DDISVFLRHI LHPSSNFVTH EMQYSSSLPH LLPKNQHGLD 60 ISNGELSSVL NSSAGGIFSS SYGAYNVPAN VSSSSVGTMD NEPDEYECES EGGTEDLGVE 120 ASTQPPPPSN TNSKRSRAAE VHNLSEKRRR SRINEKMKAL QNLIPNSNKT DKASMLDEAI 180 EYLKQLQLQV QMLTMRNGLS MYPLGLPRVL QQNQVSQLSM GLCEGNGCTN AKAAGALHVS 240 QDTLLNAIFS PSENRTETKL TPVETMSNVN HSNNAFESES SINVHLDPFQ FSRSTSKGIL 300 REDWLPLHEM SERQSKTVSI GENLASSVPF DASELKRNTL EACLLRYQSG AMSETNMDCD 360 QLLAPQLYCH AGRSTSVDTI KTECSNF |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00035 | PBM | Transfer from AT4G36930 | Download |
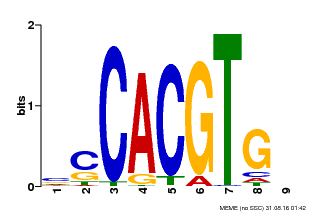
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | FJ004235 | 2e-69 | FJ004235.1 Catharanthus roseus bHLH transcription factor MYC5 (MYC5) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009783822.1 | 0.0 | PREDICTED: transcription factor SPATULA-like isoform X1 | ||||
| Refseq | XP_016464627.1 | 0.0 | PREDICTED: transcription factor SPATULA-like isoform X1 | ||||
| TrEMBL | A0A1S3ZJV8 | 0.0 | A0A1S3ZJV8_TOBAC; transcription factor SPATULA-like isoform X1 | ||||
| TrEMBL | A0A1U7WV93 | 0.0 | A0A1U7WV93_NICSY; transcription factor SPATULA-like isoform X1 | ||||
| STRING | XP_009783822.1 | 0.0 | (Nicotiana sylvestris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA9850 | 22 | 27 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G36930.1 | 2e-43 | bHLH family protein | ||||



