 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_016451536.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Nicotianoideae; Nicotianeae; Nicotiana
|
||||||||
| Family | BBR-BPC | ||||||||
| Protein Properties | Length: 333aa MW: 37283.9 Da PI: 9.0215 | ||||||||
| Description | BBR-BPC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GAGA_bind | 367.6 | 2.3e-112 | 1 | 333 | 1 | 301 |
GAGA_bind 1 mdddgsre..rnkg....yyepa.aslkenlglqlmssiaerdakirernlalsekkaavaerdmaflqrdkalaernkalverdnkllalllve 88
mdd+g++e r+k+ ++ ++ +s+k q+m+++ erda+i+ernlalsek+aa+aerdma lqrd+a+aern a++erdn++++l+++e
XP_016451536.1 1 MDDSGNHEnaRHKPpqgqWFMQHqPSMK-----QIMAIMGERDAAIQERNLALSEKRAALAERDMAILQRDSAIAERNSAIMERDNAIATLQYRE 90
7888776668888643344555556676.....************************************************************** PP
GAGA_bind 89 nsla....salpvgvqvlsgtksidslqq.lse.pqledsavelreeekle.alpieeaaeeakekkkkkkrqrakkpkekkak.kkkkkseksk 175
ns++ s +p+g+q+ +++k++++ q +++ pql++++++ ++++++e + p++ aa+e ++++++k++++ak+ ++k++ k+ kk +k+
XP_016451536.1 91 NSMNsgnmSPCPPGCQIAHEVKHMHHPQLhVHHqPQLGEPTFNHSDMHMSEsSIPLSPAAPELTKSRRNKRSKEAKEVTSSKRTsKPSKKVKKEG 185
**99999889******************964446*********9999987625788888888888887776666655444443335555455554 PP
GAGA_bind 176 kkvkkesader......................skaekksidlvlngvslDestlPvPvCsCtGalrqCYkWGnGGWqSaCCtttiSvyPLPvst 248
++ +k de+ +k+++k+++ +ln+vs+Dest+PvPvCsCtG lr YkWGnGGWqS+CCt ++S+yPLP+ +
XP_016451536.1 186 EDLNKTMLDESqewngaqemgggnddvnrqlgvTKTDWKDQGRGLNQVSFDESTMPVPVCSCTGDLRPSYKWGNGGWQSSCCTNNLSMYPLPMLP 280
5544444442257889******************************************************************************* PP
GAGA_bind 249 krrgaRiagrKmSqgafkklLekLaaeGydlsnpvDLkdhWAkHGtnkfvtir 301
++r+aRi+grKmS++af+klL++LaaeG+dlsnpvDLkd WAkHGtn+++ti+
XP_016451536.1 281 NKRHARIGGRKMSGSAFTKLLSRLAAEGHDLSNPVDLKDNWAKHGTNRYITIK 333
****************************************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01226 | 2.0E-154 | 1 | 333 | IPR010409 | GAGA-binding transcriptional activator |
| Pfam | PF06217 | 1.4E-97 | 1 | 333 | IPR010409 | GAGA-binding transcriptional activator |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 333 aa Download sequence Send to blast |
MDDSGNHENA RHKPPQGQWF MQHQPSMKQI MAIMGERDAA IQERNLALSE KRAALAERDM 60 AILQRDSAIA ERNSAIMERD NAIATLQYRE NSMNSGNMSP CPPGCQIAHE VKHMHHPQLH 120 VHHQPQLGEP TFNHSDMHMS ESSIPLSPAA PELTKSRRNK RSKEAKEVTS SKRTSKPSKK 180 VKKEGEDLNK TMLDESQEWN GAQEMGGGND DVNRQLGVTK TDWKDQGRGL NQVSFDESTM 240 PVPVCSCTGD LRPSYKWGNG GWQSSCCTNN LSMYPLPMLP NKRHARIGGR KMSGSAFTKL 300 LSRLAAEGHD LSNPVDLKDN WAKHGTNRYI TIK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional regulator that specifically binds to GA-rich elements (GAGA-repeats) present in regulatory sequences of genes involved in developmental processes. {ECO:0000269|PubMed:14731261}. | |||||
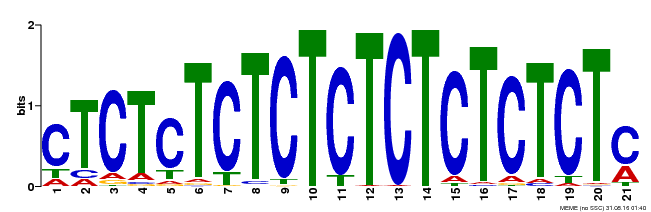
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00540 | DAP | Transfer from AT5G42520 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | FJ982302 | 0.0 | FJ982302.1 Nicotiana tabacum GAGA-binding transcriptional activator (BBR/BPC3) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001312138.1 | 0.0 | protein BASIC PENTACYSTEINE6-like | ||||
| Refseq | XP_016451534.1 | 0.0 | PREDICTED: protein BASIC PENTACYSTEINE6-like | ||||
| Refseq | XP_016451535.1 | 0.0 | PREDICTED: protein BASIC PENTACYSTEINE6-like | ||||
| Refseq | XP_016451536.1 | 0.0 | PREDICTED: protein BASIC PENTACYSTEINE6-like | ||||
| Refseq | XP_016451537.1 | 0.0 | PREDICTED: protein BASIC PENTACYSTEINE6-like | ||||
| Refseq | XP_016451538.1 | 0.0 | PREDICTED: protein BASIC PENTACYSTEINE6-like | ||||
| Refseq | XP_016451539.1 | 0.0 | PREDICTED: protein BASIC PENTACYSTEINE6-like | ||||
| Refseq | XP_016451540.1 | 0.0 | PREDICTED: protein BASIC PENTACYSTEINE6-like | ||||
| Swissprot | Q8L999 | 1e-134 | BPC6_ARATH; Protein BASIC PENTACYSTEINE6 | ||||
| TrEMBL | H1ZN85 | 0.0 | H1ZN85_TOBAC; GAGA-binding transcriptional activator | ||||
| STRING | XP_009803495.1 | 0.0 | (Nicotiana sylvestris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G42520.1 | 1e-127 | basic pentacysteine 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Entrez Gene | 107776190 |



