 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_009796558.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Nicotianoideae; Nicotianeae; Nicotiana
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 1001aa MW: 111140 Da PI: 6.0986 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 129.9 | 9.2e-41 | 145 | 222 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cqv++C +dls+ak+yhrrhkvCe+hska+ +lv +++qrfCqqCsrfh+l+efDe+krsCrrrLa+hn+rrrk+q+
XP_009796558.1 145 VCQVDDCGTDLSKAKDYHRRHKVCEMHSKASRALVGNVMQRFCQQCSRFHALQEFDEGKRSCRRRLAGHNKRRRKTQS 222
6**************************************************************************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 6.4E-33 | 140 | 207 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.086 | 143 | 220 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 7.98E-38 | 144 | 225 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 7.8E-30 | 146 | 219 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF48403 | 4.32E-7 | 773 | 886 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 4.2E-5 | 775 | 887 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 3.06E-6 | 776 | 886 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1001 aa Download sequence Send to blast |
MEASVGERYY HMGGPDLRGL GKRSLEWDVT DWKWDGDLFI ATPLRQNPSN NYQSRQFFPV 60 ETGNLVASSN SSSSCSDETN HGIEQQRREL EKRRRVIIVD ENDSGGTLSL KLGGQAEPVA 120 EKRTKLAAAA PAPAPAPVTG TTRAVCQVDD CGTDLSKAKD YHRRHKVCEM HSKASRALVG 180 NVMQRFCQQC SRFHALQEFD EGKRSCRRRL AGHNKRRRKT QSESVANNNS SNDGQASGYS 240 LMSLLKMLSN MHSNGTNHSE DQDLLAHLLR SIAGQGSLNG DKSLSGLLQE SSDLLNNRSI 300 LRNPELASLI SNGSQAPPRA KEHQFTNSAA EVPQKRLDAH DVRLEDARTA SSQSPGILFP 360 PIQSNSQAYA QGRGSTTGRS KLIDFDLNDV YVDSDDNIED IDRSPVQCPS WLQQDSHQSS 420 PPQTSGNSDS ASAQSPSSSS GDTQTRTDRI VFKLFGKDPS DFPFVVRAQI LDWLSHSPTE 480 IESYIRPGCV VLTVYLRLPE SAWEELCYDL NSSLSRLLDV HGDDSFWTKG WIYISVQNQI 540 AFVCDGQVLL DMSLPSGSNE HSTILSVRPI AVPVSGRAQF LVKGYNLSKP STRLLCALES 600 NYLVPEANNE VEEHVDGIDK DDKLQSLDFT CSVPAVTGRG FIEVEDHGLS NSFFPFIVAE 660 EDVCSEIRML ESELDLTSAN YVKGQINNME ARNQAMDFIH ELGWLLHRNN LKARLEHFGP 720 DAVLYPLKRF KWLIDFCVDH EWCAVVKKLL NVLLGGTVGA GESSFLKFAL TEMGLLHRAV 780 RRNSRPLVEL LLTYTPEKVA DELSSEYQSL VEADGEFLFR PDCVGPAGLT PLHVAAGIDG 840 SEDVLDALTD DPGKVAIEAW KNTRDSTGFT PEDYARLRGH YSYIHLVQRK ISKKAISGHI 900 VVDIPIVPSI ENSNQKEEEF ATNSLEISMT ERRPISRPCG LCHKKLAYGS RSRSLLYRPA 960 MFSMVAMAAV CVCVALLFRG SPEVLYIFRP FRWEMVDFGT S |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 7e-31 | 145 | 219 | 10 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 91 | 95 | KRRRV |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000269|PubMed:16554053}. | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' of AP1 promoter. Binds specifically to the 5'-GTAC-3' core sequence. {ECO:0000269|PubMed:10524240, ECO:0000269|PubMed:16095614}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00633 | PBM | Transfer from PK06791.1 | Download |
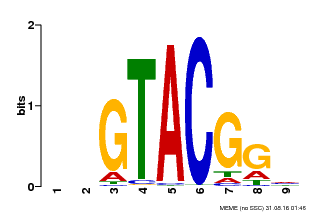
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | HG975440 | 0.0 | HG975440.1 Solanum pennellii chromosome ch01, complete genome. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009796558.1 | 0.0 | PREDICTED: squamosa promoter-binding-like protein 1 | ||||
| Swissprot | Q9S7P5 | 0.0 | SPL12_ARATH; Squamosa promoter-binding-like protein 12 | ||||
| Swissprot | Q9SMX9 | 0.0 | SPL1_ARATH; Squamosa promoter-binding-like protein 1 | ||||
| TrEMBL | A0A1U7Y0A3 | 0.0 | A0A1U7Y0A3_NICSY; squamosa promoter-binding-like protein 1 | ||||
| STRING | XP_009796558.1 | 0.0 | (Nicotiana sylvestris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA1871 | 24 | 66 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



