 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_009791697.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Nicotianoideae; Nicotianeae; Nicotiana
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 409aa MW: 44784.2 Da PI: 9.555 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 34.6 | 4.1e-11 | 331 | 386 | 5 | 59 |
CHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH.HHHHHHHHHHH CS
bZIP_1 5 krerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkke.leelkkevakl 59
+r +r++kNRe+A rsR RK+a++ eLe +v++L++ ++L+k+ +e ++k+ +l
XP_009791697.1 331 RRRKRMIKNRESAARSRDRKQAYTLELEAEVAKLKEIKQELQKKqAEFIEKQKNQL 386
689************************************99765145555555555 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 2.7E-11 | 327 | 392 | IPR004827 | Basic-leucine zipper domain |
| Gene3D | G3DSA:1.20.5.170 | 4.6E-12 | 329 | 386 | No hit | No description |
| PROSITE profile | PS50217 | 10.151 | 329 | 374 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 3.59E-9 | 331 | 379 | No hit | No description |
| CDD | cd14707 | 1.88E-22 | 331 | 385 | No hit | No description |
| Pfam | PF00170 | 2.6E-9 | 331 | 389 | IPR004827 | Basic-leucine zipper domain |
| PROSITE pattern | PS00036 | 0 | 334 | 349 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009738 | Biological Process | abscisic acid-activated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0000976 | Molecular Function | transcription regulatory region sequence-specific DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 409 aa Download sequence Send to blast |
MGSYLNFKNF ADTSQPESSG GKPMNNDNFS LARQSSIYSF TFDELQSTCG LGKDFGSMNM 60 DDLLKNIWTA EESQAMSSSV AGGNVSVPVG NLQRQGSLTL PRTISQKTVD EVWKDFQKET 120 VNANDGSAPG ASNFGQRQST LGEMTLEEFL VRAGAVREDM QPTRYSKDVT FTSGFTQPSS 180 NNSSLNIAFQ QATQNPQQLS NQIAENSIFN VVTTTSSQQK PQQAQPLFPK QTTVAFASPM 240 QLGNTAQLAS PGTRAPIVGM SNPSVNTSII QGSIMQGGVM DMAGLHNGVT SVKGGSPGNL 300 DPSSLSPSPY ACGEVGRGRR SCTSFEKVVE RRRKRMIKNR ESAARSRDRK QAYTLELEAE 360 VAKLKEIKQE LQKKQAEFIE KQKNQLLEKM NMPWENKLIC LRRTATGPW |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Binds to the ABA-responsive element (ABRE). Mediates stress-responsive ABA signaling. {ECO:0000269|PubMed:11884679, ECO:0000269|PubMed:15361142}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00038 | PBM | Transfer from AT4G34000 | Download |
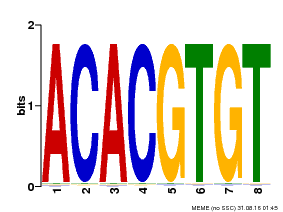
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by drought, salt, abscisic acid (ABA). {ECO:0000269|PubMed:10636868, ECO:0000269|PubMed:16284313}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB063648 | 0.0 | AB063648.1 Nicotiana tabacum mRNA for phi-2, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009791697.1 | 0.0 | PREDICTED: ABSCISIC ACID-INSENSITIVE 5-like protein 7 | ||||
| Refseq | XP_009791704.1 | 0.0 | PREDICTED: ABSCISIC ACID-INSENSITIVE 5-like protein 7 | ||||
| Swissprot | Q9M7Q3 | 3e-99 | AI5L6_ARATH; ABSCISIC ACID-INSENSITIVE 5-like protein 6 | ||||
| TrEMBL | A0A1U7XI11 | 0.0 | A0A1U7XI11_NICSY; ABSCISIC ACID-INSENSITIVE 5-like protein 7 | ||||
| STRING | XP_009791697.1 | 0.0 | (Nicotiana sylvestris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA776 | 24 | 97 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G34000.2 | 2e-85 | abscisic acid responsive elements-binding factor 3 | ||||



