 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_009790908.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Nicotianoideae; Nicotianeae; Nicotiana
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 474aa MW: 51617.9 Da PI: 8.5429 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 53.3 | 6.3e-17 | 55 | 99 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r rWT+eE++++++a+k++G W++I +++g ++t+ q++s+ qk+
XP_009790908.1 55 RERWTEEEHKKFLEAIKLYGRA-WRRIEEHVG-TKTAVQIRSHAQKF 99
78******************88.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.35E-16 | 49 | 103 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 21.896 | 50 | 104 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 4.9E-17 | 53 | 102 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 1.4E-13 | 54 | 102 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 5.9E-10 | 55 | 95 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 2.8E-14 | 55 | 98 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 7.78E-11 | 57 | 100 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0010600 | Biological Process | regulation of auxin biosynthetic process | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 474 aa Download sequence Send to blast |
MASEEMVDQD QGGESGMDSI RSSRNGISSD VGASSTRSIQ LKEQARKPYT ISKQRERWTE 60 EEHKKFLEAI KLYGRAWRRI EEHVGTKTAV QIRSHAQKFF SKVVRESNNG DASSVKPIEI 120 PPPRPKRKPM HPYPRKLATP VKSGTLVPEK PTRSISPNMS MSEQKNQSPA SVLSAHGSDA 180 LGTVDSSKRN GSASPVSPAG GGNSDGFVLS EPPNFILQDS SSLAQACVKL ELLPQEVNFV 240 KEGSAETSST PCLKLFGKTV LVTESQRPRP TSGTCKIETD VNDEAALPTA SLNLMPMKFT 300 ASDSDCGSST LTVGTPPPFY YSPSQNENQW PTGCASATLL PWGSSCASVS FPCIQVLNPI 360 PIKGRPLFND NELEDKQNQK EGSSTGSNTE PVSAEMGGDK NMDIEAQSSQ HPLEKARKEL 420 SSRPSETPSL VRRASSTKRV KGFVPYKRCL AERGTNPSML SCEEREEQRT RLCL |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00515 | DAP | Transfer from AT5G17300 | Download |
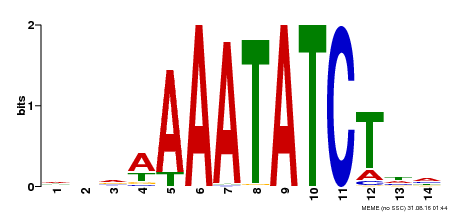
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009790908.1 | 0.0 | PREDICTED: protein REVEILLE 1-like isoform X2 | ||||
| TrEMBL | A0A1U7XJ54 | 0.0 | A0A1U7XJ54_NICSY; protein REVEILLE 1-like isoform X2 | ||||
| STRING | XP_009790907.1 | 0.0 | (Nicotiana sylvestris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G17300.1 | 3e-50 | MYB_related family protein | ||||



