 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_009788853.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Nicotianoideae; Nicotianeae; Nicotiana
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 398aa MW: 43693.2 Da PI: 7.4798 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 102.8 | 2.1e-32 | 222 | 277 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k+r++W+peLH+rF++a++qLGGs++AtPk+i+elmkv+gLt+++vkSHLQkYRl+
XP_009788853.1 222 KQRRCWSPELHRRFLQALQQLGGSHVATPKQIRELMKVDGLTNDEVKSHLQKYRLH 277
79****************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 14.865 | 219 | 279 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.24E-18 | 220 | 280 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 8.9E-29 | 220 | 280 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 7.1E-28 | 222 | 277 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 8.0E-8 | 224 | 275 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 398 aa Download sequence Send to blast |
MMINNHENNY TEKMQRCQQY IDALEQERNK IQVFSRELPL CLELVTQAIE TYKQQLSGTT 60 TEYNVHTQSD VECSEEHTSS DVPILEEFIP LKGTFSHEDE DEDERDSHKS KTYNISDTSC 120 SKDNKNSDKS CKKSDWLRSV QLWNNNQTPD PTPKEEVTPK KGAVVEVKKN GSGGAFHPFK 180 KEKSAAATTE PPTSAAVAAA AASSTAETGS GSKKEEKDEK RKQRRCWSPE LHRRFLQALQ 240 QLGGSHVATP KQIRELMKVD GLTNDEVKSH LQKYRLHTRR PSPSSIHNNH QQPPQFVVVG 300 GIWVPPPEYS AMAAAAPSAS GEASGVANSN GIYAPIATLP KGPLHDVSGG TLKQRQHNNK 360 PSRSSERGSG SHSDGGGVHS NSPATSSSTH TTTASPPY |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 6e-13 | 222 | 275 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 6e-13 | 222 | 275 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 6e-13 | 222 | 275 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 6e-13 | 222 | 275 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 6e-13 | 222 | 275 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 6e-13 | 222 | 275 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 6e-13 | 222 | 275 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 6e-13 | 222 | 275 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in phosphate signaling in roots. {ECO:0000250|UniProtKB:Q9FX67}. | |||||
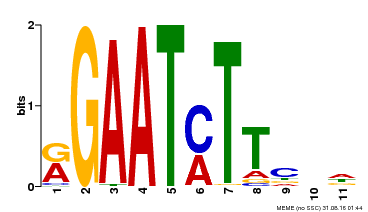
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00160 | DAP | Transfer from AT1G25550 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | HG975444 | 9e-47 | HG975444.1 Solanum pennellii chromosome ch05, complete genome. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009788853.1 | 0.0 | PREDICTED: transcription factor LUX-like | ||||
| Swissprot | Q9FPE8 | 1e-101 | HHO3_ARATH; Transcription factor HHO3 | ||||
| TrEMBL | A0A1U7XDA3 | 0.0 | A0A1U7XDA3_NICSY; transcription factor LUX-like | ||||
| STRING | XP_009788853.1 | 0.0 | (Nicotiana sylvestris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA5219 | 24 | 38 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G25550.1 | 1e-78 | G2-like family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



