 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_009782444.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Nicotianoideae; Nicotianeae; Nicotiana
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 758aa MW: 83179.5 Da PI: 6.0831 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 66 | 5.2e-21 | 55 | 110 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe+lF+++++p++++r eL+k+l+L++rqVk+WFqNrR+++k
XP_009782444.1 55 KKRYHRHTPQQIQELESLFKECPHPDEKQRLELSKRLSLETRQVKFWFQNRRTQMK 110
688999***********************************************999 PP
| |||||||
| 2 | START | 201.2 | 4.3e-63 | 262 | 487 | 1 | 206 |
HHHHHHHHHHHHHHHHC-TT-EEEE....EXCCTTEEEEEEESSS......SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S....EEEEE CS
START 1 elaeeaaqelvkkalaeepgWvkss....esengdevlqkfeeskv.....dsgealrasgvvdmvlallveellddkeqWdetla....kaetl 82
ela++a++elvk+a+ +ep+W++s+ e++n +e++++f++ + +ea+r++g+v+ ++ lve+l+d++ +W e+++ + +t+
XP_009782444.1 262 ELALAAMDELVKMAQTDEPLWFRSVeggrEILNHEEYMRSFTPCIGmrpnsLVSEASRETGMVIINSLALVETLMDSN-KWAEMFPcliaRTSTT 355
5899***************************************999****9***************************.**************** PP
EEECTT......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--..-TTSEE-EESSEEEEEEEECTCEEEEEEEE- CS
START 83 evissg......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe.sssvvRaellpSgiliepksnghskvtwveh 169
+vissg galqlm aelq+lsplvp R++ f+R+++q+ +g+w++vdvS+d ++ ++ + + ++ lpSg+l+++++ng+skvtwveh
XP_009782444.1 356 DVISSGmggtrnGALQLMHAELQVLSPLVPiREVNFLRFCKQHAEGVWAVVDVSIDTIRETSSnAPTFPNSRILPSGCLVQDMPNGYSKVTWVEH 450
*********************************************************999999999***************************** PP
EE--SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXXX CS
START 170 vdlkgrlphwllrslvksglaegaktwvatlqrqcek 206
+++++++h l+r+l++ g+ +ga++wvatlqrqce+
XP_009782444.1 451 GEYDESVIHHLYRPLISAGMGFGAQRWVATLQRQCEC 487
***********************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 2.5E-21 | 41 | 106 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 1.34E-20 | 41 | 112 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.637 | 52 | 112 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 5.2E-19 | 53 | 116 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 4.62E-19 | 54 | 112 | No hit | No description |
| Pfam | PF00046 | 1.5E-18 | 55 | 110 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 87 | 110 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 43.103 | 253 | 490 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 3.75E-32 | 255 | 487 | No hit | No description |
| CDD | cd08875 | 1.40E-122 | 257 | 486 | No hit | No description |
| SMART | SM00234 | 1.8E-47 | 262 | 487 | IPR002913 | START domain |
| Pfam | PF01852 | 7.6E-55 | 262 | 487 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 2.47E-23 | 506 | 744 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 758 aa Download sequence Send to blast |
MEGQSEITRM AENYEGNNSN GRRSREEEPD SRSGSDNLEG ASGDDQDAAD KPPRKKRYHR 60 HTPQQIQELE SLFKECPHPD EKQRLELSKR LSLETRQVKF WFQNRRTQMK TQLERHENSI 120 LRQENDKLRA ENMSIREAMR NPICTNCGGP AMIGEISLEE QHLRIENARL KDELDRVCAL 180 AGKFLGRPIS SLVNAMPPPM PNSSLELGVG NNGFGGLSNV PTTLPLAPPD FGVGISNSLP 240 VVASTRQTTG IERSLERSMY LELALAAMDE LVKMAQTDEP LWFRSVEGGR EILNHEEYMR 300 SFTPCIGMRP NSLVSEASRE TGMVIINSLA LVETLMDSNK WAEMFPCLIA RTSTTDVISS 360 GMGGTRNGAL QLMHAELQVL SPLVPIREVN FLRFCKQHAE GVWAVVDVSI DTIRETSSNA 420 PTFPNSRILP SGCLVQDMPN GYSKVTWVEH GEYDESVIHH LYRPLISAGM GFGAQRWVAT 480 LQRQCECLAI LMSSTVSSRD HTAITPSGRR SMLKLAQRMT NNFCAGVCAS TVHKWNKLCA 540 GNVDEDVRVM TRKSVDDPGE PPGIVLSAAT SVWLPISPQR LFDFLRDERL RSEWDILSNG 600 GPMQEMAHIA KGQDHGNCVS LLRASAINAN QSSMLILQET CIDAAGALVV YAPVDIPAMH 660 VVMNGGDSAY VALLPSGFSI VPDGPGSRGS NTANCGSNGP SCNGGPDQRI SGSLLTVAFQ 720 ILVNSLPTAK LTVESVETVN NLISCTVQKI KAALHCES |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
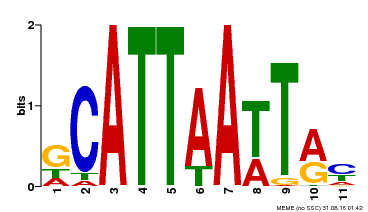
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | GQ222185 | 0.0 | GQ222185.1 Solanum lycopersicum cultivar M82 cutin deficient 2 (CD2) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009782443.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ANTHOCYANINLESS 2 isoform X1 | ||||
| Refseq | XP_016465399.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ANTHOCYANINLESS 2-like | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | A0A1S3ZM71 | 0.0 | A0A1S3ZM71_TOBAC; homeobox-leucine zipper protein ANTHOCYANINLESS 2-like | ||||
| TrEMBL | A0A1U7WV07 | 0.0 | A0A1U7WV07_NICSY; homeobox-leucine zipper protein ANTHOCYANINLESS 2 isoform X1 | ||||
| STRING | XP_009782443.1 | 0.0 | (Nicotiana sylvestris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00730.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



