 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_009768904.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Nicotianoideae; Nicotianeae; Nicotiana
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 450aa MW: 49296.2 Da PI: 5.5719 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 51.8 | 1.9e-16 | 37 | 81 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r +WT+eE++++v+a k++G W+ I +++g ++t+ q++s+ qk+
XP_009768904.1 37 REKWTEEEHQRFVEALKLYGRA-WRQIEEHVG-TKTAIQIRSHAQKF 81
789*****************88.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 5.68E-16 | 31 | 85 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 21.971 | 32 | 86 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 9.3E-17 | 35 | 84 | IPR006447 | Myb domain, plants |
| Gene3D | G3DSA:1.10.10.60 | 8.8E-9 | 36 | 77 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.0E-12 | 36 | 84 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.4E-13 | 37 | 80 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.00E-9 | 39 | 82 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 450 aa Download sequence Send to blast |
MISSKLETVA TQGKALTCLA NEITLKVRKP YTITKQREKW TEEEHQRFVE ALKLYGRAWR 60 QIEEHVGTKT AIQIRSHAQK FFAKVARDSG TDGDESLNAI DIPPPRPKKK PLHPYPRKMA 120 DSPVTNKSVS GQPERSSPPN ASGRDSCSPD SVLSEIGSDV SEYLVAEQQS SRFSPYSCTT 180 DAHAANIISG QNGDESMTSK SNIVKEIPVA SKPVAASSSS IFNSGMECDF VPRETSFPGE 240 KLAVEAPLAS IKLFGQMVLV PDATKLALQS PAGSCNSLPS KTTENEIETN NGNVIHGFQA 300 NSQFILPVVP GNMIPSACWL SQDMLENNPE GVAAFPTTIP WLSWYQDLVY RYISLCSQTA 360 AETTTLCRRP EDEEPQREGS SSGLSVGSAS EVDDGNRSSE TVESKYTTKS DKWNNIKGFV 420 PYKRCLAERD GMSSGVELEE RESQRVRVCS |
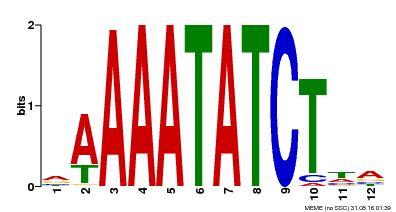
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00146 | DAP | Transfer from AT1G18330 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009768903.1 | 0.0 | PREDICTED: uncharacterized protein LOC104219851 isoform X1 | ||||
| Refseq | XP_016493565.1 | 0.0 | PREDICTED: uncharacterized protein LOC107812899 isoform X1 | ||||
| TrEMBL | A0A1S4BXE4 | 0.0 | A0A1S4BXE4_TOBAC; uncharacterized protein LOC107812899 isoform X1 | ||||
| TrEMBL | A0A1U7VR70 | 0.0 | A0A1U7VR70_NICSY; uncharacterized protein LOC104219851 isoform X1 | ||||
| STRING | XP_009768903.1 | 0.0 | (Nicotiana sylvestris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G18330.1 | 3e-38 | MYB_related family protein | ||||



