 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_009760089.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Nicotianoideae; Nicotianeae; Nicotiana
|
||||||||
| Family | TALE | ||||||||
| Protein Properties | Length: 362aa MW: 41208 Da PI: 6.2373 | ||||||||
| Description | TALE family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 28.6 | 2.4e-09 | 289 | 323 | 21 | 55 |
HSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHH CS
Homeobox 21 knrypsaeereeLAkklgLterqVkvWFqNrRake 55
k +yps++e+ LA+ +gL+++q+ +WF N+R ++
XP_009760089.1 289 KWPYPSESEKVALAETTGLDQKQINNWFINQRKRH 323
669*****************************885 PP
| |||||||
| 2 | ELK | 37.9 | 3.7e-13 | 243 | 264 | 1 | 22 |
ELK 1 ELKhqLlrKYsgyLgsLkqEFs 22
ELK++LlrKYsgyL+sLkqE+s
XP_009760089.1 243 ELKNHLLRKYSGYLSSLKQELS 264
9*******************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01255 | 4.0E-20 | 99 | 143 | IPR005540 | KNOX1 |
| Pfam | PF03790 | 1.7E-22 | 100 | 142 | IPR005540 | KNOX1 |
| SMART | SM01256 | 1.3E-25 | 150 | 201 | IPR005541 | KNOX2 |
| Pfam | PF03791 | 1.5E-23 | 154 | 199 | IPR005541 | KNOX2 |
| Pfam | PF03789 | 2.5E-10 | 243 | 264 | IPR005539 | ELK domain |
| PROSITE profile | PS51213 | 11.296 | 243 | 263 | IPR005539 | ELK domain |
| SMART | SM01188 | 3.5E-7 | 243 | 264 | IPR005539 | ELK domain |
| PROSITE profile | PS50071 | 12.924 | 263 | 326 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 1.67E-19 | 264 | 338 | IPR009057 | Homeodomain-like |
| SMART | SM00389 | 5.4E-13 | 265 | 330 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 2.7E-27 | 268 | 329 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 5.44E-12 | 275 | 327 | No hit | No description |
| Pfam | PF05920 | 8.5E-17 | 283 | 322 | IPR008422 | Homeobox KN domain |
| PROSITE pattern | PS00027 | 0 | 301 | 324 | IPR017970 | Homeobox, conserved site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0001708 | Biological Process | cell fate specification | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010051 | Biological Process | xylem and phloem pattern formation | ||||
| GO:0010089 | Biological Process | xylem development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 362 aa Download sequence Send to blast |
MENNYNQLGE NSAQRGNNFL YGGIPLLASP NSSIYGRSSG GGGDCYDQSQ AVVHPIVKTE 60 GGSTSHHHTF QYPSIIRSGY HHLTVQNHHE SESSGSEVDA LKAKIIAHPQ CSNLLDAYMD 120 CQKVGAPPEV VARLSAVRQE FEVRQRDSST DRNIAKDPEL DQFMEAYYDM LVKYREELTR 180 PLHEAMDFMR KIETQLNMLG NGPVRIFNSE DNCEGVGSSE EEQDNSGGET EIPEIDPRAE 240 DRELKNHLLR KYSGYLSSLK QELSKKKKKG KLPKDARQKL LSWWELHYKW PYPSESEKVA 300 LAETTGLDQK QINNWFINQR KRHWKPSEDM QFMVMDGLHP QNAAALYMEG HYMGEGPFRL 360 GQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Appears to be involved in meristem formation and in the regulation of leaf morphology. Misexpression makes the leaf more compound which is always associated with growth retardation and loss of apical dominance, resulting in dwarfed, bushy plants. Probably binds to the DNA sequence 5'-TGAC-3'. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00675 | ChIP-seq | Transfer from GRMZM2G017087 | Download |
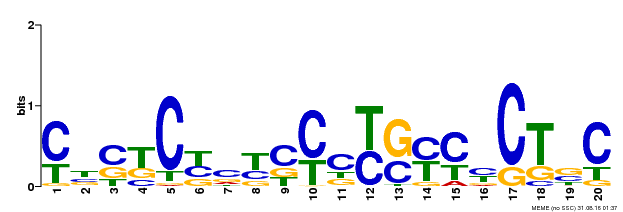
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AF544054 | 0.0 | AF544054.1 Nicotiana tabacum knotted 3 (kn3) mRNA, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009760089.1 | 0.0 | PREDICTED: homeotic protein knotted-1 isoform X1 | ||||
| Refseq | XP_016442876.1 | 0.0 | PREDICTED: homeotic protein knotted-1-like isoform X1 | ||||
| Swissprot | Q41330 | 0.0 | KN1_SOLLC; Homeotic protein knotted-1 | ||||
| TrEMBL | A0A1S3XSE7 | 0.0 | A0A1S3XSE7_TOBAC; homeotic protein knotted-1-like isoform X1 | ||||
| TrEMBL | A0A1U7VCR6 | 0.0 | A0A1U7VCR6_NICSY; homeotic protein knotted-1 isoform X1 | ||||
| STRING | XP_009760089.1 | 0.0 | (Nicotiana sylvestris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA4973 | 23 | 33 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G08150.1 | 1e-137 | KNOTTED-like from Arabidopsis thaliana | ||||



