 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | NNU_013977-RA | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; stem eudicotyledons; Proteales; Nelumbonaceae; Nelumbo
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 827aa MW: 90813 Da PI: 6.5907 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 58.5 | 1.1e-18 | 19 | 77 | 3 | 57 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHC....TS-HHHHHHHHHHHHHHHHC CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkkl....gLterqVkvWFqNrRakekk 57
k ++t+eq+e+Le+l++++++ps +r++L +++ +++ +q+kvWFqNrR +ek+
NNU_013977-RA 19 KYVRYTPEQVEALERLYHECPKPSSIRRQQLIRECpilsNIEPKQIKVWFQNRRCREKQ 77
5679*****************************************************97 PP
| |||||||
| 2 | START | 180.9 | 7.5e-57 | 163 | 371 | 2 | 205 |
HHHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-SEEEEEEEECTT..EEEEEEE CS
START 2 laeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeqWdetlakaetlevissg..galqlmv 95
+aee+++e+++ka+ ++ Wv+++ +++g++++ +++ s++++g a+ra+g+v +++ v+e+l+d++ W ++++++++l+v+ +g g+++l +
NNU_013977-RA 163 IAEETLTEFLSKATGTAVEWVQMPGMKPGPDSIGIVAISHGCTGVAARACGLVGLEPT-RVAEILKDRPSWFRDCRAVDVLNVLPTGngGTIELLY 257
789*******************************************************.8888888888*************************** PP
EXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--....-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SSXXHHHHHHHHHH CS
START 96 aelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe...sssvvRaellpSgiliepksnghskvtwvehvdlkgrlphwllrslvks 187
++l+a+++l+p Rdf+ +Ry+ l++g++v++++S++++q p+ +++vR+e+lpSg+li+p+++g+s +++v+h+dl+ ++++++lr+l++s
NNU_013977-RA 258 MQLYAPTTLAPaRDFWLLRYTSVLEDGSLVVCERSLSNTQGGPSmptVQNFVRGEMLPSGYLIRPCEGGGSIIHIVDHMDLEPWSVPEVLRPLYES 353
******************************************9988889*********************************************** PP
HHHHHHHHHHHHTXXXXX CS
START 188 glaegaktwvatlqrqce 205
+++ ++kt++a+l+++++
NNU_013977-RA 354 STVLAQKTTMAALRHLRQ 371
*************99876 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 9.6E-19 | 8 | 77 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 15.758 | 14 | 78 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 2.8E-15 | 16 | 82 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 9.11E-17 | 19 | 79 | No hit | No description |
| SuperFamily | SSF46689 | 4.71E-17 | 19 | 82 | IPR009057 | Homeodomain-like |
| Pfam | PF00046 | 2.9E-16 | 20 | 77 | IPR001356 | Homeobox domain |
| CDD | cd14686 | 1.13E-6 | 71 | 110 | No hit | No description |
| PROSITE profile | PS50848 | 26 | 153 | 381 | IPR002913 | START domain |
| CDD | cd08875 | 1.56E-81 | 157 | 373 | No hit | No description |
| SuperFamily | SSF55961 | 1.65E-38 | 162 | 374 | No hit | No description |
| Gene3D | G3DSA:3.30.530.20 | 2.1E-23 | 162 | 368 | IPR023393 | START-like domain |
| SMART | SM00234 | 4.1E-43 | 162 | 372 | IPR002913 | START domain |
| Pfam | PF01852 | 2.1E-54 | 163 | 371 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 6.46E-5 | 399 | 494 | No hit | No description |
| SuperFamily | SSF55961 | 6.46E-5 | 522 | 584 | No hit | No description |
| Pfam | PF08670 | 7.5E-52 | 684 | 826 | IPR013978 | MEKHLA |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0010014 | Biological Process | meristem initiation | ||||
| GO:0010075 | Biological Process | regulation of meristem growth | ||||
| GO:0010087 | Biological Process | phloem or xylem histogenesis | ||||
| GO:0048263 | Biological Process | determination of dorsal identity | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 827 aa Download sequence Send to blast |
MMAVTASCKD GKGGMENGKY VRYTPEQVEA LERLYHECPK PSSIRRQQLI RECPILSNIE 60 PKQIKVWFQN RRCREKQRKE ASRLQAVNRK LTAMNKLLME ENDRLQKQVS QLVYENGYFR 120 QQTQSTALAT TDTSCESVVT SGQHHLTPQH PPRDASPAGL LSIAEETLTE FLSKATGTAV 180 EWVQMPGMKP GPDSIGIVAI SHGCTGVAAR ACGLVGLEPT RVAEILKDRP SWFRDCRAVD 240 VLNVLPTGNG GTIELLYMQL YAPTTLAPAR DFWLLRYTSV LEDGSLVVCE RSLSNTQGGP 300 SMPTVQNFVR GEMLPSGYLI RPCEGGGSII HIVDHMDLEP WSVPEVLRPL YESSTVLAQK 360 TTMAALRHLR QIAQEVSQST VTGWGRRPAA LRALSQRLSR GFNEALNGFT DEGWSMMGND 420 GMDDVTILVN SSPSKVMGVN LTFGSGFPAM NNSVLCAKAS MLLQNVPPAI LLRFLREHRS 480 EWADSNIDAY SAASLKAGPC SLPGSRLGSF GGQFLEVIKL ESIGHCQEDP MMPRDMFLLQ 540 LCSGVDENAV GTCAELIFAP IDASFADDAP LLPSGFRIIP LDTGVDASSP NRTLDLASAL 600 EVGTAGNRIS GDYSGNTGNM RSVMTIAFQF AFESHLQENV ASMARQYVRS IISSVQRVAL 660 ALSPSRLSSH SGLRPPPGTP EAHTLARWIC HSYRNFLGVE LLKPSSEGSE SILKTLWHHQ 720 DAIMCCSLKA LPVFTFANQV GLDMLETTLV ALQDITLEKI FDDHGRKTLC SEFPQIMQQG 780 FACLQGGICL SSMGRPVSYE RAVAWKVLNE EENAHCICFL FVNWSFV |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of meristem development to promote lateral organ formation. May regulates procambial and vascular tissue formation or maintenance, and vascular development in inflorescence stems. {ECO:0000269|PubMed:15598805, ECO:0000269|PubMed:15705957, ECO:0000269|PubMed:16617092}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00200 | DAP | Transfer from AT1G52150 | Download |
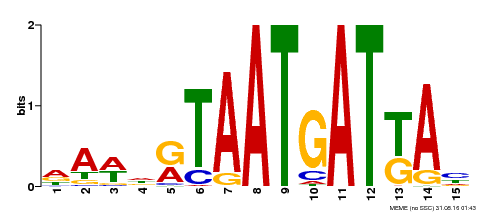
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. Repressed by miR165 and miR166. {ECO:0000269|PubMed:15773855, ECO:0000269|PubMed:16033795, ECO:0000269|PubMed:16617092, ECO:0000269|PubMed:17237362}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010267183.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ATHB-15-like | ||||
| Refseq | XP_010267184.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ATHB-15-like | ||||
| Swissprot | Q9ZU11 | 0.0 | ATB15_ARATH; Homeobox-leucine zipper protein ATHB-15 | ||||
| TrEMBL | A0A1U8AW35 | 0.0 | A0A1U8AW35_NELNU; homeobox-leucine zipper protein ATHB-15-like | ||||
| STRING | XP_010267183.1 | 0.0 | (Nelumbo nucifera) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G52150.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



