 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | NNU_009809-RA | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; stem eudicotyledons; Proteales; Nelumbonaceae; Nelumbo
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 865aa MW: 99496.3 Da PI: 7.2814 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 83.5 | 3.6e-26 | 70 | 155 | 2 | 91 |
FAR1 2 fYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkktekerrtraetrtgCkaklkvkkekdgkwevtkleleHnHelap 91
+Y+eYAk++GF +++++s++sk +ge++++++ Cs++g+++e++k ++r+ ++t+Cka+++vk+++dgkw+++++ ++HnHel p
NNU_009809-RA 70 CYKEYAKSMGFGISIKNSRRSKISGEFIDAKYACSRYGSKHESSKV----INPRPCSKTDCKASMHVKRRQDGKWVIHNFIKDHNHELLP 155
7*****************************************9888....9************************************975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 3.9E-24 | 70 | 155 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 2.0E-33 | 268 | 361 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 8.951 | 548 | 584 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 9.1E-5 | 549 | 580 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 8.4E-5 | 559 | 586 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010018 | Biological Process | far-red light signaling pathway | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 865 aa Download sequence Send to blast |
MAIDLEQPSG EHEAENVGPN EGGITVDDGE IHSGEVDVNS SMVESIVKGG ANAEPHIGME 60 FESKGAAYSC YKEYAKSMGF GISIKNSRRS KISGEFIDAK YACSRYGSKH ESSKVINPRP 120 CSKTDCKASM HVKRRQDGKW VIHNFIKDHN HELLPSQAHF FRSHRNINID ALHAIKRRRK 180 MYALMSKQPD GNQDFGYPNK DIINEFDKGR YLVLEGGDMQ AMLEHFMDMQ DENSNFFYAM 240 DLNEEQRLRN VFWVDAKGRQ DYINFGDVVC FDTTYISSKY KMPLVPFIGV NHHYQFTLFG 300 FALIADETTS TFLWLMNTWL RSMGGRAPNV IVTDQDKALK AAIAEIFPNT RHCFCLWHIL 360 RKIPEKLGHI TKRHENFMKK FNKCIYRSCT EEQFEKRWWK MVDRFELKTD EWFQSLYEDR 420 KYWVPYYMND TFFAGMSTAQ RSESINSSLD RYIHRKTTLK EFVEHYVVVL QDRYEEEAKA 480 DLDTWHKQPA LRSPSPFEKQ MSTIYTHAIF RKFQVEVLGV VACHPKMEKE DETTITYRVQ 540 DFEEQQDFLV SWNERKLEIS CLCHSYEYKG FLCRHAMIVL QISGVSNIPS HYVLKRWTKD 600 AKSRHTMRQG LEGVQSRVQR YNDLCQRAIK LGEEGSLSQD SYNIAFRALD EALRECVGVN 660 NSIQSAVNAN ISATYGIHDI EEEDQGNNTM SMLRMLDAHA TRTSKKKNAN KKRKIHLEPK 720 VTNSGIENSL QDMGQLRAPT LDGSFIAQQS IQGMEQLSSR APAFDTYGTQ QTMQRLGRLN 780 SVMATRDSYY GNQQGMQGLG QLHSIAPSSD SYYGTQQSMQ AIGQIDFRSP TIDGCYGLQD 840 SLPDMGESAQ LHGVARHLHD KHLSP |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
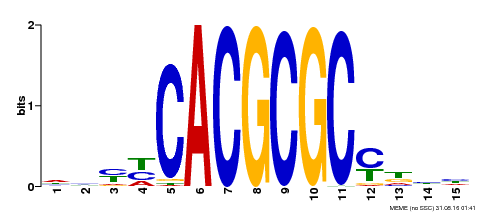
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00434 | DAP | Transfer from AT4G15090 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. {ECO:0000269|PubMed:18033885}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010260557.1 | 0.0 | PREDICTED: protein FAR-RED IMPAIRED RESPONSE 1 isoform X1 | ||||
| Swissprot | Q9SWG3 | 0.0 | FAR1_ARATH; Protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| TrEMBL | A0A1U8AE84 | 0.0 | A0A1U8AE84_NELNU; protein FAR-RED IMPAIRED RESPONSE 1 isoform X1 | ||||
| STRING | XP_010260557.1 | 0.0 | (Nelumbo nucifera) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G15090.1 | 0.0 | FAR1 family protein | ||||



