 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | NNU_005712-RA | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; stem eudicotyledons; Proteales; Nelumbonaceae; Nelumbo
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 701aa MW: 76797.7 Da PI: 6.917 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 75.3 | 7.2e-24 | 122 | 223 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT.......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrse 89
f k+lt+sd+++ g +++p+ +ae++ ++ + t+ +d +g++W++++iyr++++r++lt+GW+ Fv++++L +gD++vF + ++
NNU_005712-RA 122 FAKTLTQSDANNGGGFSVPRYCAETIfprldysADPPVQ--TVLAKDVHGEVWKFRHIYRGTPRRHLLTTGWSTFVNQKKLIAGDSIVFL--RAEN 213
89****************************996555555..8999*********************************************..4499 PP
E..EEEEE-S CS
B3 90 felvvkvfrk 99
+el+v+++r+
NNU_005712-RA 214 GELCVGIRRA 223
99******97 PP
| |||||||
| 2 | Auxin_resp | 118.9 | 4.1e-39 | 291 | 374 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrl.sGtvvgvsdldpvrWpnSkWrsLk 83
aa+ a++++ FevvY+Prast+eF+vk++ v++a+++++++GmRfkmafetedss++++ +Gt+++v+ +dp+rWpnS+Wr+L+
NNU_005712-RA 291 AATLAANGQSFEVVYYPRASTPEFCVKASAVKAAMRIQWCPGMRFKMAFETEDSSRISWfMGTISSVQVADPIRWPNSPWRLLQ 374
68899*****************************************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.40.330.10 | 8.3E-37 | 113 | 226 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 2.88E-28 | 120 | 228 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 2.21E-21 | 121 | 222 | No hit | No description |
| Pfam | PF02362 | 5.0E-22 | 122 | 223 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 2.7E-22 | 122 | 224 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 13.084 | 122 | 224 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 3.6E-34 | 291 | 374 | IPR010525 | Auxin response factor |
| PROSITE profile | PS51745 | 19.971 | 615 | 695 | IPR000270 | PB1 domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007389 | Biological Process | pattern specification process | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0048829 | Biological Process | root cap development | ||||
| GO:0051301 | Biological Process | cell division | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 701 aa Download sequence Send to blast |
MITFTDSVKE PMKETEKCLD SQLWHACAGG MVQMPSINSK VFYFPQGQAE HAHGNVDFGN 60 SPRIPPLILC RVAAVKYLAD PETDEVYAKI RLVPLRNNEP DYDDDGVLGS NGSDVAEKPA 120 SFAKTLTQSD ANNGGGFSVP RYCAETIFPR LDYSADPPVQ TVLAKDVHGE VWKFRHIYRG 180 TPRRHLLTTG WSTFVNQKKL IAGDSIVFLR AENGELCVGI RRAKRGVGGG PESLSGWNPA 240 AGNCVSPYGG FSVFLREDEN KLMRNVNGGN SNSGVGLRTR GRVRAESVIE AATLAANGQS 300 FEVVYYPRAS TPEFCVKASA VKAAMRIQWC PGMRFKMAFE TEDSSRISWF MGTISSVQVA 360 DPIRWPNSPW RLLQVTWDEP DLLQNVKCVS PWLVELVSNM PAIHLSPFSP PRKKFRLPQH 420 PDFPLDSQFQ IPSFSGNPLG PSSPLCCLSD NTPAGIQGAR HAQFGISLSD LHLNKLQSGL 480 FPASFQRLDN TTPPSKINNG LIRGNPNGSE NISCLLTIGS STQSSRKSDG RKSPGFLLFG 540 QPILTEQQIS LSCSGETVSP VLTGNSSSEG NNQDKATNFS DGSGSALNQL GPLDNSCEGF 600 PWYKERPTTE IGLETGHCKV FMESEDVGRT LDLTVLGSYE ELYRRLANMF GIERSEMLSR 660 VLYQDATGAV KHTGDEPFSE FMKTARRLTI LMDSGSDNIG R |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldu_A | 2e-77 | 19 | 395 | 51 | 388 | Auxin response factor 5 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
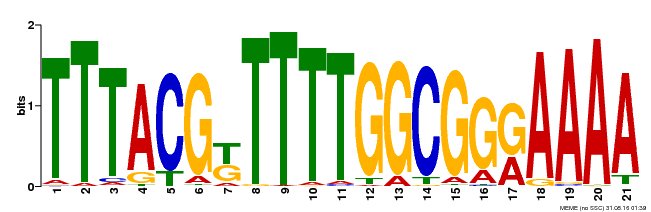
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00461 | DAP | Transfer from AT4G30080 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010253698.1 | 0.0 | PREDICTED: auxin response factor 18-like | ||||
| Swissprot | Q653H7 | 0.0 | ARFR_ORYSJ; Auxin response factor 18 | ||||
| TrEMBL | A0A1U7ZKL2 | 0.0 | A0A1U7ZKL2_NELNU; Auxin response factor | ||||
| STRING | XP_010253698.1 | 0.0 | (Nelumbo nucifera) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G30080.1 | 0.0 | auxin response factor 16 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



