 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | NNU_004056-RA | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; stem eudicotyledons; Proteales; Nelumbonaceae; Nelumbo
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 395aa MW: 43523 Da PI: 7.8317 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 131.6 | 2.6e-41 | 92 | 168 | 2 | 78 |
-SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 2 CqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
C v+gC++dls++++yhrrhkvCe hsk+p+v+v g+eqrfCqqCsrfh l efDe krsCr+rL++hn+rrrk+q+
NNU_004056-RA 92 CLVDGCKSDLSNCRDYHRRHKVCELHSKTPKVMVGGQEQRFCQQCSRFHLLVEFDEVKRSCRKRLDGHNRRRRKPQP 168
**************************************************************************997 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 4.5E-34 | 85 | 153 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 31.992 | 89 | 166 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 4.19E-39 | 90 | 171 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 2.8E-30 | 92 | 165 | IPR004333 | Transcription factor, SBP-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0048653 | Biological Process | anther development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 395 aa Download sequence Send to blast |
MDWDLKTPSW DLAELEQEAI PNIAPLVGSS SFEGQKTKED CSVDLKLGRL GDSGDGSLDK 60 LKDPRIATML SSPSGSTKRA RTPGNGTQTV SCLVDGCKSD LSNCRDYHRR HKVCELHSKT 120 PKVMVGGQEQ RFCQQCSRFH LLVEFDEVKR SCRKRLDGHN RRRRKPQPES LAMNSGSLFP 180 KHQGTRLMPF ATTEMFPTTV VSPAWSAVVK TEVDATRYNH QQQLNFIDKQ NTFPGSFSRS 240 CYRGGKHFPF LQSNDVTLIN RTPSEASMCQ PLLNSIASSE SEGGRKMFSN GLTRVIDSDC 300 ALSLLSSPPM QSSGISMNHV VQPDPIPMAQ PLVPVLQYNG IGRYPCSQGM EDEQGGSVLA 360 PDASNADLHC HGVFQLGPYG SDRNGGPQTL PFSWE |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 6e-27 | 92 | 165 | 11 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 148 | 165 | KRSCRKRLDGHNRRRRKP |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
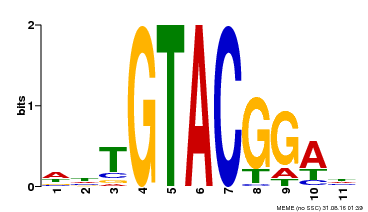
| Motif ID | Method | Source | Motif file |
| MP00555 | DAP | Transfer from AT5G50570 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010250660.1 | 0.0 | PREDICTED: teosinte glume architecture 1-like | ||||
| Refseq | XP_010250661.1 | 0.0 | PREDICTED: teosinte glume architecture 1-like | ||||
| Refseq | XP_010250662.1 | 0.0 | PREDICTED: teosinte glume architecture 1-like | ||||
| TrEMBL | A0A1U7ZAX4 | 0.0 | A0A1U7ZAX4_NELNU; teosinte glume architecture 1-like | ||||
| STRING | XP_010250660.1 | 0.0 | (Nelumbo nucifera) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G50670.1 | 3e-60 | SBP family protein | ||||



