 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | NNU_001887-RA | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; stem eudicotyledons; Proteales; Nelumbonaceae; Nelumbo
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1048aa MW: 117402 Da PI: 6.442 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 69.2 | 6.6e-22 | 36 | 100 | 2 | 74 |
CG-1 2 lkekkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLK 74
l+ ++rwl++ ei++iL n+ k++++ e++++p+s ++ryfrkDG++w+kkkdgkt++E+he+LK
NNU_001887-RA 36 LEAQNRWLRPAEICEILRNYPKFRIAPEPPNKPPSM--------VLRYFRKDGHNWRKKKDGKTIKEAHERLK 100
5679*****************************974........67**************************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 33.414 | 31 | 157 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 5.6E-17 | 34 | 117 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 2.4E-17 | 37 | 102 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 3.9E-6 | 462 | 549 | IPR013783 | Immunoglobulin-like fold |
| Pfam | PF01833 | 1.1E-7 | 463 | 548 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 2.18E-18 | 463 | 549 | IPR014756 | Immunoglobulin E-set |
| CDD | cd00204 | 1.99E-13 | 643 | 755 | No hit | No description |
| SuperFamily | SSF48403 | 2.64E-16 | 649 | 758 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 17.661 | 650 | 767 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 3.6E-17 | 650 | 759 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF00023 | 0.015 | 696 | 727 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.0058 | 696 | 725 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 9.19 | 696 | 728 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 190 | 735 | 764 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 1.38E-8 | 865 | 922 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.083 | 871 | 893 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.444 | 872 | 901 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0019 | 873 | 892 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0046 | 894 | 916 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.597 | 895 | 919 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 2.6E-4 | 897 | 916 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1048 aa Download sequence Send to blast |
MLCVYQPQEN KHSVISQIKV QNSTYISRTD IEQILLEAQN RWLRPAEICE ILRNYPKFRI 60 APEPPNKPPS MVLRYFRKDG HNWRKKKDGK TIKEAHERLK GAKTNFGRMR DTEEVVPSSQ 120 MGSPMSSSFL TNNTQVPSQT MDTTSLNSTQ ASEYEDAESD NHQASSRYHS IFESQQSEDS 180 AVMNKMDANL LNSYYPDPCQ NNYQGKKPAV PGLDFVSLVQ ENRGRDGNDA RFLPTSEPQK 240 QVNLTCWDVL EHCTTGFQNA SFQPLILSSQ PAAIGVIPKE ESVIPGQFLA EEFTNPEIAG 300 QPDGQEKWQT ASVDNSSYMS RWPKDQKLHP DPAYALEAFH MHPDQQNGHP VQNDLPIQIS 360 GAELASVLKS NSDHNLTMEG NPYNAKQPIE FSQTEEGLKK LDSFTRWMTK ELGEVDESHT 420 KLSSVDWNAV ENGTEVDNSG MSQAHLHSYL LSPSISQDQL FSIIDFSPNW AYTDSEVKVL 480 ITGTFLRTQE DAAKCKWSCM FGEVEVLAEV IGDGVLRCHA PPHTAGRVPF YVTRSNRLAC 540 SEVREFEYRV KHTRMDATNM SSSTNEILLH VRLGKLLSMG CSSHPTTLTS NVGEKAHISN 600 KISLLMKEDD DEWFHMVKLI LEEFSPDQIK DQLLQKLLKE KLHAWLLYKV IEDGKGPSVL 660 DKEGQGVLHL SAALGYDWAI APTVAAGVSI NFRDVNGWTA LHWAAFYGRE RTVVALVTLG 720 AAPGALTDPT PKFPSGRTPA DLASSNGHKG IAGYLAEAAL TSHLSSITLK DTKDGDAPEI 780 SGMKAVQTVS ERSATPGCDG DVLDGLSLKD SLTAVRNATQ AAARIHQVFR VQSFQRKQLT 840 EYGDNKFGMS DEHALSLLSV KTHRAGQHDD PLHSAAIRIQ NKFRGWKGRK EFLIIRQRIV 900 KIQAHVRGHQ VRKRYKNIVW SVGIVEKAIL RWRRKGSGLR GFRPEPLIEG SSTQNDPSKE 960 DDYDFLKEGR KQTEERLQKA LARVKSMAQY PEARDQYRRL LNVVSEFQDA KVMYDKVLNG 1020 SEEAGEGDDD LIELEALLDD DTFMATAC |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
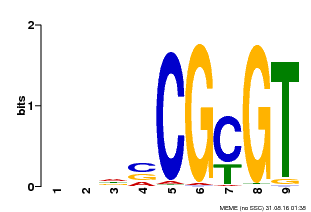
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010267840.1 | 0.0 | PREDICTED: calmodulin-binding transcription activator 3-like | ||||
| TrEMBL | A0A1U8AKK4 | 0.0 | A0A1U8AKK4_NELNU; calmodulin-binding transcription activator 3-like | ||||
| STRING | XP_010267832.1 | 0.0 | (Nelumbo nucifera) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 0.0 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||



