 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | NNU_001193-RA | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; stem eudicotyledons; Proteales; Nelumbonaceae; Nelumbo
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 366aa MW: 40772.4 Da PI: 7.2772 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 103.2 | 1.6e-32 | 203 | 258 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
kpr++W+peLH+rF++a++qLGGs+ AtPk+i+elmkv+gLt+++vkSHLQkYRl+
NNU_001193-RA 203 KPRRCWSPELHRRFLHALQQLGGSHAATPKQIRELMKVDGLTNDEVKSHLQKYRLH 258
79****************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 14.047 | 200 | 260 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.23E-18 | 201 | 261 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 2.4E-29 | 201 | 261 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 5.3E-27 | 203 | 258 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.4E-7 | 205 | 256 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 366 aa Download sequence Send to blast |
MNPQPTMTIM ANYTENAHKC QTYMEALEEE RRKIQVFRRE LPLCLELVTQ AIETCRQQLA 60 TTTEYFPGQS ECSEETSSEA PVLEEFIPLK RCFSSEEDEE SNKPKNEDIN EKPGKKADWL 120 KSVQLWNQDP DPSLNQDLSR KPSVIESKKG GGAFHPFQRD DKLTAPSITV IPSSAQTTAA 180 ISSTADTING DNRGEDKSQS NRKPRRCWSP ELHRRFLHAL QQLGGSHAAT PKQIRELMKV 240 DGLTNDEVKS HLQKYRLHTR RPSPAVQNNG SPQPPQFVVV GGIWVPPHDY AAATETKGEL 300 NGVSGVTAHP TIYAPITAPS VPTVRSEQRQ KLSPRSPLQA EGRSSCKKDD DTHSNSPSTS 360 SSSRSY |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 2e-13 | 203 | 256 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 2e-13 | 203 | 256 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_A | 3e-13 | 203 | 256 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 3e-13 | 203 | 256 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 3e-13 | 203 | 256 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 3e-13 | 203 | 256 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 2e-13 | 203 | 256 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 2e-13 | 203 | 256 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 2e-13 | 203 | 256 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 2e-13 | 203 | 256 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 2e-13 | 203 | 256 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 2e-13 | 203 | 256 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in phosphate signaling in roots. {ECO:0000250|UniProtKB:Q9FX67}. | |||||
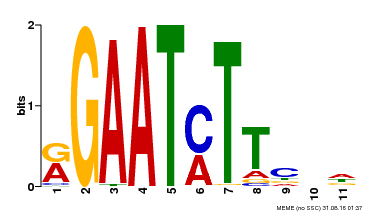
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00160 | DAP | Transfer from AT1G25550 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010263063.1 | 0.0 | PREDICTED: myb family transcription factor EFM-like isoform X2 | ||||
| Swissprot | Q9FPE8 | 1e-107 | HHO3_ARATH; Transcription factor HHO3 | ||||
| TrEMBL | A0A1U8A8T7 | 0.0 | A0A1U8A8T7_NELNU; myb family transcription factor EFM-like isoform X2 | ||||
| STRING | XP_010263055.1 | 0.0 | (Nelumbo nucifera) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G25550.1 | 1e-105 | G2-like family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



