 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Niben101Scf09590g03004.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Nicotianoideae; Nicotianeae; Nicotiana
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 467aa MW: 51035 Da PI: 8.6212 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 50.5 | 4.8e-16 | 44 | 88 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r rW+ eE+ ++++a k++G W++I +++g ++t+ q++s+ qk+
Niben101Scf09590g03004.1 44 RERWSDEEHRKFIEALKLHGRA-WRRIEEHVG-TKTAVQIRSHAQKF 88
78******************88.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 8.22E-16 | 38 | 92 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 21.61 | 39 | 93 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 3.8E-15 | 42 | 91 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 5.9E-13 | 43 | 91 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.8E-13 | 44 | 87 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 7.2E-9 | 44 | 84 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 5.28E-10 | 46 | 89 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0010600 | Biological Process | regulation of auxin biosynthetic process | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 467 aa Download sequence Send to blast |
MTIQDQSNSD TILPRGSRTS LDVGAPADEY APKIRKPYTI SKQRERWSDE EHRKFIEALK 60 LHGRAWRRIE EHVGTKTAVQ IRSHAQKFFS KVVRESNNGD VSSVKSIEIP PPRPKRKPMH 120 PYPRKLAIPV KSGTLAPEIS KRSASPNLCL SETENQSPTS VLSALGSDAF GTVDSSKPSE 180 SPSPVSSAVS ENCGDLVLSE QSDFNLKKRR SSPAQAYASS NPDNKACVKL ELFPEDNDFV 240 KEGSVEASST HSLKLFGKTV FVTDSCVLSS PTSGQILLAD VNDEPASRTL SQSSFPMKFS 300 PSDSECAPNT VTMYHVPAAN GSPRLNKSVS CDPLSWGSSY VSAAFSCIQV HNPIPMKGRA 360 LFDHKDSENQ ETQKEGSSTG SNTESVSAEL SVDKSLGVEA QISRNQEAGR EFFSIRASAP 420 SVFERRANST KRVTGFVPYK RCLGERGISS STLTGEEREE KRTRLCL |
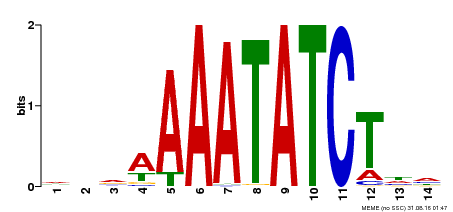
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00515 | DAP | Transfer from AT5G17300 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009782662.1 | 0.0 | PREDICTED: protein REVEILLE 1-like isoform X1 | ||||
| TrEMBL | A0A1U7WRV3 | 0.0 | A0A1U7WRV3_NICSY; protein REVEILLE 1-like isoform X1 | ||||
| STRING | XP_009782662.1 | 0.0 | (Nicotiana sylvestris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA3185 | 24 | 47 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G17300.1 | 8e-57 | MYB_related family protein | ||||



