 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Niben101Scf08693g00002.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Nicotianoideae; Nicotianeae; Nicotiana
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 841aa MW: 92371.8 Da PI: 6.5419 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 58.9 | 8.1e-19 | 17 | 75 | 3 | 57 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHC....TS-HHHHHHHHHHHHHHHHC CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkkl....gLterqVkvWFqNrRakekk 57
k ++t+eq+e+Le+l++ +++ps +r++L +++ +++ +q+kvWFqNrR +ek+
Niben101Scf08693g00002.1 17 KYVRYTPEQVEALERLYHDCPKPSSMRRQQLIRECpilsNIEPKQIKVWFQNRRCREKQ 75
5679*****************************************************97 PP
| |||||||
| 2 | START | 174.6 | 6e-55 | 161 | 369 | 2 | 205 |
HHHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-SEEEEEEEEC CS
START 2 laeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeqWdetlakaetlevis 86
+aee+++e+++ka+ ++ Wv+++ +++g++++ +++ s++++g a+ra+g+v +++ v+e+l+d++ W ++++++++l+v+
Niben101Scf08693g00002.1 161 IAEETLTEFLSKATGTAVEWVQMPGMKPGPDSIGIIAISHGCTGMAARACGLVGLEPT-RVAEILKDRPSWFRDCRAVDVLNVLP 244
789*******************************************************.8888888888**************** PP
TT..EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--....-TTSEE-EESSEEEEEEEECTCEEEEE CS
START 87 sg..galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe...sssvvRaellpSgiliepksnghskvt 165
++ g+++l +++l+a+++l+p Rdf+ +Ry+ + +g++v++++S+ ++q+ p+ +++vRae lpSg+li+p+++g+s ++
Niben101Scf08693g00002.1 245 TAngGTIELLYMQLYAPTTLAPaRDFWLLRYTTVMDDGSLVVCERSLGNTQNGPSmppVQNFVRAEILPSGYLIRPCEGGGSIIH 329
9999************************************************99988899************************* PP
EEE-EE--SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXX CS
START 166 wvehvdlkgrlphwllrslvksglaegaktwvatlqrqce 205
+v+h++l++++++++lr+l++s+++ ++kt+va+l+++++
Niben101Scf08693g00002.1 330 IVDHMNLESWSVPEVLRPLYESSAVLAQKTTVAALRYLRQ 369
***********************************99876 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50071 | 15.516 | 12 | 76 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 5.2E-16 | 14 | 80 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 8.02E-17 | 17 | 77 | No hit | No description |
| SuperFamily | SSF46689 | 8.98E-17 | 17 | 80 | IPR009057 | Homeodomain-like |
| Pfam | PF00046 | 2.2E-16 | 18 | 75 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 2.1E-18 | 19 | 75 | IPR009057 | Homeodomain-like |
| CDD | cd14686 | 1.15E-6 | 69 | 108 | No hit | No description |
| PROSITE profile | PS50848 | 26.074 | 151 | 379 | IPR002913 | START domain |
| CDD | cd08875 | 4.63E-76 | 155 | 371 | No hit | No description |
| SMART | SM00234 | 1.1E-42 | 160 | 370 | IPR002913 | START domain |
| Gene3D | G3DSA:3.30.530.20 | 2.1E-22 | 160 | 365 | IPR023393 | START-like domain |
| SuperFamily | SSF55961 | 3.71E-37 | 160 | 372 | No hit | No description |
| Pfam | PF01852 | 1.1E-52 | 161 | 369 | IPR002913 | START domain |
| Pfam | PF08670 | 5.6E-50 | 698 | 840 | IPR013978 | MEKHLA |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0010014 | Biological Process | meristem initiation | ||||
| GO:0010075 | Biological Process | regulation of meristem growth | ||||
| GO:0010087 | Biological Process | phloem or xylem histogenesis | ||||
| GO:0048263 | Biological Process | determination of dorsal identity | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 841 aa Download sequence Send to blast |
MASCKDGKTS SVMDNGKYVR YTPEQVEALE RLYHDCPKPS SMRRQQLIRE CPILSNIEPK 60 QIKVWFQNRR CREKQRKEAS RLQAVNRKLT AMNKLLMEEN DRLQKQVSQL VYENGYFRRQ 120 THSTTLATKD TSCESVVTSG QHHLTSQHPP RDASPAGLLS IAEETLTEFL SKATGTAVEW 180 VQMPGMKPGP DSIGIIAISH GCTGMAARAC GLVGLEPTRV AEILKDRPSW FRDCRAVDVL 240 NVLPTANGGT IELLYMQLYA PTTLAPARDF WLLRYTTVMD DGSLVVCERS LGNTQNGPSM 300 PPVQNFVRAE ILPSGYLIRP CEGGGSIIHI VDHMNLESWS VPEVLRPLYE SSAVLAQKTT 360 VAALRYLRQI AQEVSQTNVT NWGRRPAALR ALSQRLSRGF NEALNGLTDE GWSMLGCDGM 420 DDITILVNSS PDKLMGLNLS FANGFSSLSN AVMCAKASML LQNVPPAILL RFLREHRSEW 480 ADNNIDAYSA AAIKVGPCCL PGARVGNLGG QVILPLAHTV EHEEASLLLE VIKLEGVGHS 540 PEDAIMPRDM FLLQLCSGMD ENAVGTSAEL IFAPIDASFA DDAPLLPSGF RIISLESGKE 600 SSSPNRTLDL TSALETGPAE NKVGNDLHTN GGASRSVMTI AFQFAFESHM QENVASMARQ 660 YVRSIISSVQ RVALALSPSH LGSHGGLRLP LGTPEAHTLA RWISQSYRCF LGVELLKSIA 720 EGSESILKSL WHHSDAIICC SAKAMPVFTF ANQAGLDMLE TTLVALQDIT LEKIFDDHGK 780 KNLCTEFPQI MQQGFACLQG GICLSSMSRP ISYERAVAWK VTNEEDSPHC ICFMFVNWSF 840 V |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of meristem development to promote lateral organ formation. May regulates procambial and vascular tissue formation or maintenance, and vascular development in inflorescence stems. {ECO:0000269|PubMed:15598805, ECO:0000269|PubMed:15705957, ECO:0000269|PubMed:16617092}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
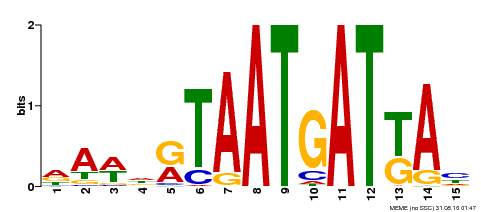
| Motif ID | Method | Source | Motif file |
| MP00200 | DAP | Transfer from AT1G52150 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. Repressed by miR165 and miR166. {ECO:0000269|PubMed:15773855, ECO:0000269|PubMed:16033795, ECO:0000269|PubMed:16617092, ECO:0000269|PubMed:17237362}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_019267274.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ATHB-15-like isoform X1 | ||||
| Swissprot | Q9ZU11 | 0.0 | ATB15_ARATH; Homeobox-leucine zipper protein ATHB-15 | ||||
| TrEMBL | A0A314KZL5 | 0.0 | A0A314KZL5_NICAT; Homeobox-leucine zipper protein athb-15 | ||||
| STRING | XP_009774805.1 | 0.0 | (Nicotiana sylvestris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA457 | 24 | 140 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G52150.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



