 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Niben101Scf04372g00012.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Nicotianoideae; Nicotianeae; Nicotiana
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 845aa MW: 92184.1 Da PI: 5.9637 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 65.8 | 6e-21 | 128 | 183 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe+lF+++++p++++r eL+k+l+L++rqVk+WFqNrR+++k
Niben101Scf04372g00012.1 128 KKRYHRHTPQQIQELESLFKECPHPDEKQRLELSKRLSLETRQVKFWFQNRRTQMK 183
688999***********************************************999 PP
| |||||||
| 2 | START | 192.5 | 2e-60 | 335 | 574 | 1 | 206 |
HHHHHHHHHHHHHHHHC-TT-EEEE...................EXCCTTEEEEEEESSS......SCEEEEEEEECCSCHHHHH CS
START 1 elaeeaaqelvkkalaeepgWvkss...................esengdevlqkfeeskv.....dsgealrasgvvdmvlall 61
ela++a++elvk+a+ +ep+W++s+ e++n +e+++++++ + + +ea+r++g+v+ ++ l
Niben101Scf04372g00012.1 335 ELALAAMDELVKMAQTDEPLWFRSVeggreilnheexrsveggrEILNHEEYMRSLTPCIGmrpnsFVSEASREIGMVIINSLAL 419
5899*************************99999999999999999999999999999888****9******************* PP
HHHHCCCGGCT-TT-S....EEEEEEEECTT......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS CS
START 62 veellddkeqWdetla....kaetlevissg......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqk 135
ve+l+d++ +W e+++ + +t++vissg galqlm aelq+lsplvp R++ f+R+++q+ +g+w++vdvS+d ++
Niben101Scf04372g00012.1 420 VETLMDSN-KWAEMFPcliaRTSTTDVISSGmggtrnGALQLMHAELQVLSPLVPiREVNFLRFCKQHAEGVWAVVDVSIDTIRE 503
********.**************************************************************************** PP
--.-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXXX CS
START 136 ppesssvvRaellpSgiliepksnghskvtwvehvdlkgrlphwllrslvksglaegaktwvatlqrqcek 206
++ + + ++ lpSg+l+++++ng+skv+wveh +++++++h l+r+l++ g+ +ga++wvatlqrqce+
Niben101Scf04372g00012.1 504 TSNAPTFPNSRILPSGCLVQDMPNGYSKVIWVEHGEYDENVIHHLYRPLISAGMGFGAQRWVATLQRQCEC 574
*99******************************************************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 2.8E-21 | 114 | 179 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 1.55E-20 | 114 | 185 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.637 | 125 | 185 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 5.2E-19 | 126 | 189 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 5.62E-19 | 127 | 185 | No hit | No description |
| Pfam | PF00046 | 1.7E-18 | 128 | 183 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 160 | 183 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 41.78 | 326 | 577 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 1.21E-29 | 328 | 378 | No hit | No description |
| CDD | cd08875 | 2.91E-116 | 330 | 573 | No hit | No description |
| Pfam | PF01852 | 3.5E-52 | 335 | 574 | IPR002913 | START domain |
| SMART | SM00234 | 3.5E-42 | 335 | 574 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 1.21E-29 | 405 | 574 | No hit | No description |
| SuperFamily | SSF55961 | 1.98E-23 | 593 | 831 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 845 aa Download sequence Send to blast |
MNFGDFLDNT SGGGGSGGGG ARIVADIPFN NNNSNSSNNN MPAGAISQPR LLTQSLAKSM 60 FNSPGLSLAL QTGMEGQSEV TRMAENYEGN NSNGRRSREE EPDSRSGSDN LEGASGDDQD 120 AADKPPRKKR YHRHTPQQIQ ELESLFKECP HPDEKQRLEL SKRLSLETRQ VKFWFQNRRT 180 QMKTQLERHE NSILRQENDK LRAENMSIRE AMRNPICTNC GGPAMIGEIS LEEQHLRIEN 240 ARLKDELDRV CALAGKFLGR PISSLVNSMP PPMPNSSLEL GVGNNGFGGL SNVPTTLPLA 300 PPDFGVGISN SLPVVASTRQ TTGIERSLER SMYLELALAA MDELVKMAQT DEPLWFRSVE 360 GGREILNHEE XRSVEGGREI LNHEEYMRSL TPCIGMRPNS FVSEASREIG MVIINSLALV 420 ETLMDSNKWA EMFPCLIART STTDVISSGM GGTRNGALQL MHAELQVLSP LVPIREVNFL 480 RFCKQHAEGV WAVVDVSIDT IRETSNAPTF PNSRILPSGC LVQDMPNGYS KVIWVEHGEY 540 DENVIHHLYR PLISAGMGFG AQRWVATLQR QCECLAILMS STVSSRDHTA ITPSGRRSML 600 KLAQRMTNNF CAGVCASTVH KWNKLCAGNV DEDVRVMTRK SVDDPGEPPG IVLSAATSVW 660 LPVSPQRLFD FLRDERLRSE WDILSNGGPM QEMAHIAKGQ DHGNCVSLLR ASAINANQSS 720 MLILQETCID AAGALVVYAP VDIPAMHVVM NGGDSAYVAL LPSGFSIVPD GPGSRGSTTA 780 NFGSNGPSSN GGPDQRISGS LLTVAFQILV NSLPTAKLTV ESVETVNNLI SCTVQKIKAA 840 LHCES |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
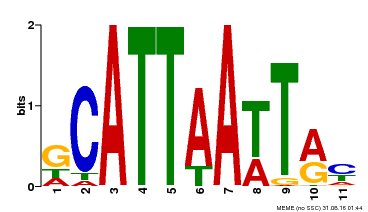
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_019259406.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | A0A1J6ISN2 | 0.0 | A0A1J6ISN2_NICAT; Homeobox-leucine zipper protein anthocyaninless 2 | ||||
| STRING | XP_009602856.1 | 0.0 | (Nicotiana tomentosiformis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA1626 | 24 | 65 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00730.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



