 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Medtr8g432540.1 | ||||||||
| Common Name | MTR_8g432540 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Trifolieae; Medicago
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 811aa MW: 88904.5 Da PI: 5.8989 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 65.6 | 6.6e-21 | 116 | 171 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe+lF+++++p++++r eL+k+l L++rqVk+WFqNrR+++k
Medtr8g432540.1 116 KKRYHRHTPQQIQELESLFKECPHPDEKQRLELSKRLCLETRQVKFWFQNRRTQMK 171
688999***********************************************999 PP
| |||||||
| 2 | START | 205.4 | 2.3e-64 | 320 | 544 | 1 | 206 |
HHHHHHHHHHHHHHHHC-TT-EEEE....EXCCTTEEEEEEESSS......SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S....EEEE CS
START 1 elaeeaaqelvkkalaeepgWvkss....esengdevlqkfeeskv.....dsgealrasgvvdmvlallveellddkeqWdetla....kaet 81
ela++a++elvk+a+ ++p+W +s e++n +e++++f++ + + +ea+r++g v+ ++ lve+l+d++ +W e+++ + +t
Medtr8g432540.1 320 ELALAAMDELVKMAQTSDPLWIRSIeggrEILNHEEYMRTFTPCIGlkpngFVSEASRETGTVIINSLALVETLMDSN-RWIEMFPciiaRTST 412
5899**************************************99999*******************************.*************** PP
EEEECTT......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--.-TTSEE-EESSEEEEEEEECTCEEEEEEEE CS
START 82 levissg......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppesssvvRaellpSgiliepksnghskvtwve 168
evis+g galqlm+ael +lsplvp R++ f+R+++q+ +g+w++vdvS+d ++ + s+v +++lpSg+++++++ng+skvtwve
Medtr8g432540.1 413 NEVISNGingtrnGALQLMQAELHVLSPLVPvREVNFLRFCKQHAEGVWAVVDVSIDTIRETSGATSFVNCRKLPSGCVVQDMPNGYSKVTWVE 506
*************************************************************999****************************** PP
-EE--SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXXX CS
START 169 hvdlkgrlphwllrslvksglaegaktwvatlqrqcek 206
h++++++++h+l+r+l++sg+ +ga++wvatlqrqce+
Medtr8g432540.1 507 HTEYEENQVHQLYRPLLSSGMGFGASRWVATLQRQCEC 544
************************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.5E-20 | 101 | 173 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 1.1E-21 | 105 | 173 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.248 | 113 | 173 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 2.2E-17 | 114 | 177 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 3.83E-18 | 115 | 173 | No hit | No description |
| Pfam | PF00046 | 1.9E-18 | 116 | 171 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 148 | 171 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 44.084 | 311 | 547 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 6.04E-34 | 313 | 544 | No hit | No description |
| CDD | cd08875 | 2.03E-124 | 315 | 543 | No hit | No description |
| Pfam | PF01852 | 7.2E-56 | 320 | 544 | IPR002913 | START domain |
| SMART | SM00234 | 6.1E-48 | 320 | 544 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 7.03E-24 | 572 | 803 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 811 aa Download sequence Send to blast |
MSFGGFVENN SGGGGGVRNI GEISYNNERM PFGSFSQPRL VTSPALAKSM FNSPGLSLAL 60 QTNIDGQEDV NGSMHGNYEQ NGLRRSREEE QSRSGSDNLD GVSGDEQDAD DKPPRKKRYH 120 RHTPQQIQEL ESLFKECPHP DEKQRLELSK RLCLETRQVK FWFQNRRTQM KTQLERHENN 180 LLRQENDKLR AENMSIRDAM RNPICSNCGG PAMIGEISLE EQHLRIDNAR LKDELDRVCA 240 LAGKFLGRPI SSLPNSSLEL GVGGNNNNGF NVMNNVPSTL PDFSSGMSNN PLAIVSPSNR 300 QTSMVTTGFD RSVERSMFLE LALAAMDELV KMAQTSDPLW IRSIEGGREI LNHEEYMRTF 360 TPCIGLKPNG FVSEASRETG TVIINSLALV ETLMDSNRWI EMFPCIIART STNEVISNGI 420 NGTRNGALQL MQAELHVLSP LVPVREVNFL RFCKQHAEGV WAVVDVSIDT IRETSGATSF 480 VNCRKLPSGC VVQDMPNGYS KVTWVEHTEY EENQVHQLYR PLLSSGMGFG ASRWVATLQR 540 QCECLAILMS SAAPSRDHSA ITAGGRRSML KLAQRMTNNF CAGVCASTVH KWNKLNPGNV 600 DEDVRVMTRK SVDDPGEPPG IVLSAATSVW LPVSPQRLFD FLRDERLRSE WDILSNGGPM 660 QEMAHIAKGQ DHGNCVSLLR ASAMNSNQSS MLILQETCID EAGSLVVYAP VDIPAMHVVM 720 NGGDSAYVAL LPSGFAIVPD GPGSRGLENG DATANGGGEA RVSGSLLTVA FQILVNSLPT 780 AKLTVESVET VNNLISCTVQ KIKAALQCES * |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Mtr.14899 | 0.0 | leaf | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, stems, leaves and floral buds. {ECO:0000269|PubMed:10402424}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
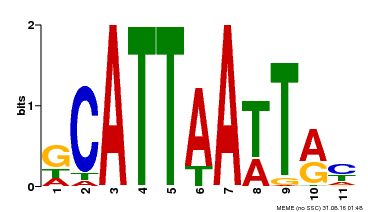
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Medtr8g432540.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_013444922.1 | 0.0 | homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | A0A072TNH2 | 0.0 | A0A072TNH2_MEDTR; Homeobox leucine zipper protein | ||||
| STRING | XP_004510857.1 | 0.0 | (Cicer arietinum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1192 | 34 | 106 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00730.1 | 0.0 | HD-ZIP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Medtr8g432540.1 |
| Entrez Gene | 25502315 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



