 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Medtr7g069660.1 | ||||||||
| Common Name | MTR_7g069660 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Trifolieae; Medicago
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 353aa MW: 40077.7 Da PI: 5.4599 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 99.9 | 1.8e-31 | 72 | 125 | 2 | 55 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
prlrWtp+LH rF++av++LGG+e+AtPk +l+lm++kgL+++hvkSHLQ+YR+
Medtr7g069660.1 72 PRLRWTPDLHFRFLHAVQRLGGQERATPKLVLQLMNIKGLSIAHVKSHLQMYRS 125
8****************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 11.179 | 68 | 128 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.7E-28 | 68 | 126 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 2.33E-14 | 72 | 126 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.1E-22 | 72 | 127 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 5.8E-9 | 73 | 124 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 353 aa Download sequence Send to blast |
MNMREEEAYG SECSKASISI NEDGCEISEG NEDEESNKQN INNGGISSSN STIEENCEKK 60 SSVRPYVRSK FPRLRWTPDL HFRFLHAVQR LGGQERATPK LVLQLMNIKG LSIAHVKSHL 120 QMYRSKKVVD TNQVLADHRL LVDNGDRNVY NLTQLPMLQG YTPNQTSSYR CGSYGDASLA 180 MYENMVQMSS ISDSRADFYG KMIERTNNNI INIQGHSIFQ MDSSNFRELS TTKVHEPNDN 240 FLSFCGHESL RDDLQVQPRA QDLISNDNLP ANQVEELKTM KRKASDMDLE LDLSLKLNSR 300 NDHKCIEEHE IDSNLSLSLY SQSSYVCKRM KEEQDGSKEQ EKGVNISLDL TI* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 3e-18 | 73 | 127 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 3e-18 | 73 | 127 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_A | 4e-18 | 73 | 127 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 4e-18 | 73 | 127 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 4e-18 | 73 | 127 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 4e-18 | 73 | 127 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 3e-18 | 73 | 127 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 3e-18 | 73 | 127 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 3e-18 | 73 | 127 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 3e-18 | 73 | 127 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 3e-18 | 73 | 127 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 3e-18 | 73 | 127 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00297 | DAP | Transfer from AT2G38300 | Download |
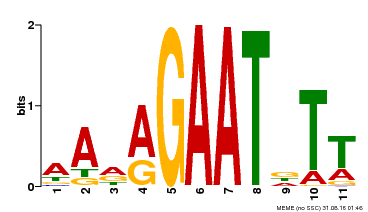
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Medtr7g069660.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC146554 | 0.0 | AC146554.5 Medicago truncatula clone mth2-9l7, complete sequence. | |||
| GenBank | CT967302 | 0.0 | CT967302.9 M.truncatula DNA sequence from clone MTH2-146M13 on chromosome 3, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003623331.2 | 0.0 | putative Myb family transcription factor At1g14600 | ||||
| TrEMBL | G7KSC8 | 0.0 | G7KSC8_MEDTR; MYB-like transcription factor family protein | ||||
| STRING | AES79549 | 0.0 | (Medicago truncatula) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF2378 | 34 | 82 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G38300.1 | 1e-54 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Medtr7g069660.1 |
| Entrez Gene | 11408041 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



