 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Medtr5g079220.1 | ||||||||
| Common Name | MTR_5g079220 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Trifolieae; Medicago
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 277aa MW: 31744.1 Da PI: 8.0507 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 52.8 | 8.9e-17 | 23 | 70 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT eEd+ll+ + +++G g W++++++ g++R++k+c++rw++yl
Medtr5g079220.1 23 KGAWTYEEDKLLKACMQKYGEGKWHLVPQRAGLNRCRKSCRLRWLNYL 70
79********************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 47.9 | 3.2e-15 | 76 | 121 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
r +++++E + +++++k+lG++ W++Ia++++ gRt++++k++w+++l
Medtr5g079220.1 76 RESFSEDEVDMILRLHKLLGNR-WSLIAARLP-GRTANDVKNYWHTHL 121
678*******************.*********.************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 4.0E-22 | 15 | 73 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 17.264 | 18 | 70 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.6E-28 | 20 | 117 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 5.4E-13 | 22 | 72 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.1E-15 | 23 | 70 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.14E-9 | 25 | 70 | No hit | No description |
| PROSITE profile | PS51294 | 24.743 | 71 | 125 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 9.0E-24 | 74 | 126 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 8.5E-16 | 75 | 123 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.7E-14 | 76 | 121 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 2.13E-11 | 78 | 121 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0031540 | Biological Process | regulation of anthocyanin biosynthetic process | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 277 aa Download sequence Send to blast |
MPLFVCGQKQ VYMEKCKTRG VRKGAWTYEE DKLLKACMQK YGEGKWHLVP QRAGLNRCRK 60 SCRLRWLNYL NPTINRESFS EDEVDMILRL HKLLGNRWSL IAARLPGRTA NDVKNYWHTH 120 LRKKMVSRKE EKKENEKPKE SMQTHEVIKP QPRTFSSHSP WLNGKYNNFV TPIVTVSTND 180 GNVAKDSEVD TILPINGDGD SAAQPYLENP TLSSMWWESL LNVSNDKIGS CSLLLPEEYS 240 KLNVENFLAE GPSTVGDFSW DSTICEFDSL LDDILN* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 2e-24 | 23 | 125 | 7 | 108 | B-MYB |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator, when associated with BHLH002/EGL3/MYC146, BHLH012/MYC1, or BHLH042/TT8. {ECO:0000269|PubMed:15361138}. | |||||
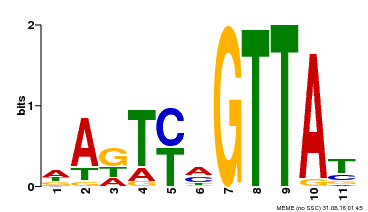
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00213 | DAP | Transfer from AT1G66370 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Medtr5g079220.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | FJ199997 | 0.0 | FJ199997.1 Medicago truncatula MYB transcription factor LAP4 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003616359.1 | 0.0 | transcription factor MYB113 | ||||
| Swissprot | Q9FNV9 | 1e-62 | MY113_ARATH; Transcription factor MYB113 | ||||
| TrEMBL | G7K1W9 | 0.0 | G7K1W9_MEDTR; Putative transcription factor MYB-HB-like family | ||||
| STRING | AES99317 | 0.0 | (Medicago truncatula) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF31 | 34 | 817 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G66370.1 | 4e-63 | myb domain protein 113 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Medtr5g079220.1 |
| Entrez Gene | 11426711 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



