 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Medtr4g121020.2 | ||||||||
| Common Name | MTR_4g121020 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Trifolieae; Medicago
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 578aa MW: 62931.1 Da PI: 6.4668 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 92.2 | 4.2e-29 | 111 | 164 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kpr++W+ eLH++Fv+av+qL G++kA+Pk+ilelm+v+gLt+e+v+SHLQkYRl
Medtr4g121020.2 111 KPRVVWSVELHQQFVDAVDQL-GIDKAVPKKILELMNVPGLTRENVASHLQKYRL 164
79*******************.********************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50110 | 15.569 | 1 | 47 | IPR001789 | Signal transduction response regulator, receiver domain |
| Gene3D | G3DSA:3.40.50.2300 | 7.9E-12 | 5 | 65 | No hit | No description |
| SuperFamily | SSF52172 | 3.42E-8 | 5 | 52 | IPR011006 | CheY-like superfamily |
| SuperFamily | SSF46689 | 3.4E-20 | 108 | 168 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 11.502 | 108 | 167 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.9E-30 | 110 | 169 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.2E-25 | 111 | 164 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 7.7E-8 | 113 | 163 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009873 | Biological Process | ethylene-activated signaling pathway | ||||
| GO:0010082 | Biological Process | regulation of root meristem growth | ||||
| GO:0010119 | Biological Process | regulation of stomatal movement | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0071368 | Biological Process | cellular response to cytokinin stimulus | ||||
| GO:0080113 | Biological Process | regulation of seed growth | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 578 aa Download sequence Send to blast |
MFCMMIIVMS ADDGKSVVMK GVTHGACDYL IKPVRIEALK NIWQHVVRKK KNEWKDTEQS 60 GSADEGDRHP KASDDADYSS SANEGNWRNS RKRRDDEEEG DDRDDSSTLK KPRVVWSVEL 120 HQQFVDAVDQ LGIDKAVPKK ILELMNVPGL TRENVASHLQ KYRLYLRRLS GVSQHQNNSF 180 LSPQEASFGT ISSMNGLDLQ TLAAAGQLPA QSLATLQAAG LGRSTVKSGL PMPLMDQRNL 240 FSFENPRLRF GEGQQQLLNG SNNKPTNLLH GVPTNMEPKQ LANLHQSAAQ SLGNMNMRVP 300 ASATQGNPLL MQMAQSQPRG PMLSENTGSH VPRLPNLLGQ STVSNGISDG LMGRNNMASS 360 SRAPSFNPVP QSSSFLNFPM NQSTEMSVSN FPLGSTPGIS SITTKGSFQA EVNSGIKGSA 420 GFPSYDIFNE LNHQKSRDWG MTNQGLMYDS SSHANPLHGN IDVSPSVLVH QGFSSAQQTG 480 HSRDASLLGK SPFSLGEGLE QGNLQNGGGQ HFNPLYVENL TRVKAERVPD ASSHTDLFPD 540 HFGQDDLMSA LLKQQEGIGQ GENEFDFDGY SLDNIPV* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 2e-22 | 110 | 170 | 4 | 64 | ARR10-B |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Mtr.9046 | 0.0 | flower| glandular trichome| leaf| root| stem | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Detected in the whole plant. Predominantly expressed in pollen. {ECO:0000269|PubMed:11370868, ECO:0000269|PubMed:15173562, ECO:0000269|PubMed:9891419}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specifically to the DNA sequence 5'-[AG]GATT-3'. Functions as a response regulator involved in His-to-Asp phosphorelay signal transduction system. Phosphorylation of the Asp residue in the receiver domain activates the ability of the protein to promote the transcription of target genes. Could directly activate some type-A response regulators in response to cytokinins. Involved in the expression of nuclear genes for components of mitochondrial complex I. Promotes cytokinin-mediated leaf longevity. Involved in the ethylene signaling pathway in an ETR1-dependent manner and in the cytokinin signaling pathway. {ECO:0000269|PubMed:11370868, ECO:0000269|PubMed:11574878, ECO:0000269|PubMed:15282545, ECO:0000269|PubMed:16407152}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00054 | PBM | Transfer from AT4G16110 | Download |
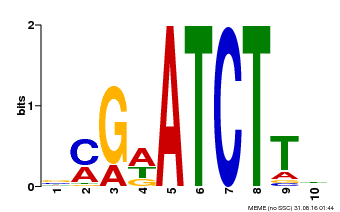
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Medtr4g121020.2 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_013458294.1 | 0.0 | two-component response regulator ARR2 isoform X1 | ||||
| Swissprot | Q9ZWJ9 | 1e-152 | ARR2_ARATH; Two-component response regulator ARR2 | ||||
| TrEMBL | A0A072USL7 | 0.0 | A0A072USL7_MEDTR; Two-component response regulator ARR2-like protein | ||||
| STRING | GLYMA17G03380.1 | 0.0 | (Glycine max) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G16110.1 | 1e-141 | response regulator 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Medtr4g121020.2 |
| Entrez Gene | 25494105 |



