 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Medtr4g100420.1 | ||||||||
| Common Name | MTR_4g100420 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Trifolieae; Medicago
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 308aa MW: 34704.5 Da PI: 5.3285 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 62.9 | 6.9e-20 | 156 | 206 | 2 | 55 |
AP2 2 gykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
+y+GVr+++ +g+++AeIrdp+++g r++lg+f ta eAaka++ a+ +l+g
Medtr4g100420.1 156 HYRGVRQRP-WGKFAAEIRDPNKRG--SRVWLGTFETAIEAAKAYDSAAFRLRG 206
69*******.**********96655..*************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00018 | 5.94E-31 | 155 | 214 | No hit | No description |
| PROSITE profile | PS51032 | 24.236 | 156 | 214 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 5.2E-14 | 156 | 206 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 3.2E-38 | 156 | 220 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 6.54E-22 | 156 | 216 | IPR016177 | DNA-binding domain |
| Gene3D | G3DSA:3.30.730.10 | 2.0E-32 | 156 | 215 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 2.2E-13 | 157 | 168 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 2.2E-13 | 196 | 216 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 308 aa Download sequence Send to blast |
MANFEEASAL KRIKMHLLGE FSPLPSPIST PCFDFDFDFE FQTIPNISQS ESNSSTSDSS 60 ISLDHYLTNL LETDTQIPLF EFDTKPQLME HESPKALTSS HHVNQPQRTV EKKPKLSRKP 120 SLEIALPKKT EWIQFGKPDP KPEVVVQKPE VAEKQHYRGV RQRPWGKFAA EIRDPNKRGS 180 RVWLGTFETA IEAAKAYDSA AFRLRGSKAI LNFPLEVNAV AASAETTVEG NKKRCREEEE 240 EEEEEVVEVK PVVKKEKITE YDMNCFKEMP LTPSFWTGLW DGDVKGLVSV PPLSPLPSFC 300 LSPLVVV* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2gcc_A | 3e-33 | 152 | 219 | 1 | 68 | ATERF1 |
| 3gcc_A | 3e-33 | 152 | 219 | 1 | 68 | ATERF1 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Mtr.6697 | 0.0 | flower| leaf| pod| root| stem | ||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Acts as a transcriptional activator. Binds to the GCC-box pathogenesis-related promoter element. Involved in the regulation of gene expression by stress factors and by components of stress signal transduction pathways. {ECO:0000269|PubMed:10715325, ECO:0000269|PubMed:9756931}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00549 | DAP | Transfer from AT5G47230 | Download |
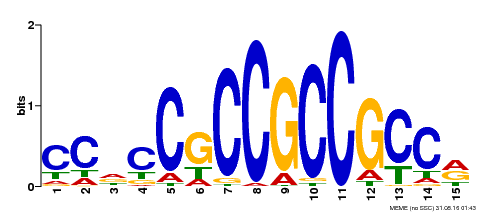
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Medtr4g100420.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Ethylene induction is completely dependent on a functional ETHYLENE-INSENSITIVE2 (EIN2). Wounding as well as cold stress induction does not require EIN2. Transcripts accumulate strongly in cycloheximide-treated plants, a protein synthesis inhibitor. Seems to not be influenced by jasmonate, Alternaria brassicicola, exogenous abscisic acid (ABA), cold, heat, NaCl or drought stress. {ECO:0000269|PubMed:10715325}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC153643 | 0.0 | AC153643.25 Medicago truncatula chromosome 8 clone mth2-109d3, complete sequence. | |||
| GenBank | BT142623 | 0.0 | BT142623.1 Medicago truncatula clone JCVI-FLMt-22N6 unknown mRNA. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003608660.2 | 0.0 | ethylene-responsive transcription factor 6 | ||||
| Swissprot | O80341 | 2e-61 | EF102_ARATH; Ethylene-responsive transcription factor 5 | ||||
| TrEMBL | G7JHN6 | 0.0 | G7JHN6_MEDTR; Ethylene response factor | ||||
| STRING | AES90857 | 0.0 | (Medicago truncatula) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1085 | 34 | 109 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G17490.1 | 1e-48 | ethylene responsive element binding factor 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Medtr4g100420.1 |
| Entrez Gene | 11405320 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



