 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Medtr3g112510.1 | ||||||||
| Common Name | MTR_3g112510 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Trifolieae; Medicago
|
||||||||
| Family | SRS | ||||||||
| Protein Properties | Length: 295aa MW: 32711.7 Da PI: 8.3563 | ||||||||
| Description | SRS family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | DUF702 | 202.8 | 9.7e-63 | 80 | 227 | 2 | 152 |
DUF702 2 rsgtasCqdCGnqakkdCaheRCRtCCksrgfdCathvkstWvpaakrrerqqqlaaasskaaasaaeaaskrkrelkskkqsalsstklssae 95
rsg+++CqdCGnqakkdC h RCRtCCksrgf+C+thvkstWvpaakrrerqq+l+a+ ++ + ++ kr+ e ++ +++l +
Medtr3g112510.1 80 RSGGMNCQDCGNQAKKDCIHLRCRTCCKSRGFQCQTHVKSTWVPAAKRRERQQHLSALPFRRRD--HVDNYKRHTEI-NQGATSLPFPQ--PPI 168
6899***************************************************998666544..55666666654.33444444444..344 PP
DUF702 96 skkeletsslPeevsseav.frcvrvssvdd.geeelaYqtavsigGhvfkGiLydqGl 152
+++ le ++P+evss+ v frcvrvss+d ++e++aYqtav+igGhvfkGiLydqG+
Medtr3g112510.1 169 PTTGLELGQFPPEVSSPGVtFRCVRVSSLDGpDHEQCAYQTAVNIGGHVFKGILYDQGP 227
556799999*******9866**********967778**********************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF05142 | 2.7E-58 | 83 | 227 | IPR007818 | Protein of unknown function DUF702 |
| TIGRFAMs | TIGR01623 | 4.5E-27 | 85 | 127 | IPR006510 | Zinc finger, lateral root primordium type 1 |
| TIGRFAMs | TIGR01624 | 9.4E-22 | 177 | 227 | IPR006511 | Lateral Root Primordium type 1, C-terminal |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009299 | Biological Process | mRNA transcription | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 295 aa Download sequence Send to blast |
MAGLFCLGRK EGQKEDKLDY YSNKELLLRN EEIYNNKGLE IWPQQSLNIY NAFGVGHSSS 60 SSSRNSSFDE CERRFGIMRR SGGMNCQDCG NQAKKDCIHL RCRTCCKSRG FQCQTHVKST 120 WVPAAKRRER QQHLSALPFR RRDHVDNYKR HTEINQGATS LPFPQPPIPT TGLELGQFPP 180 EVSSPGVTFR CVRVSSLDGP DHEQCAYQTA VNIGGHVFKG ILYDQGPPNL YTSIAPIEGS 240 SGGGHHQPQQ LDFMAATATT ATTSGIHFDP SLYSAPLNAF MAAGTQFFQP QHPN* |
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: In young plants, detected at a low level only in the root tips and lateral root primordia. In fully open flowers, primarily observed in the filaments and anthers. {ECO:0000269|PubMed:20706774}. | |||||
| Uniprot | TISSUE SPECIFICITY: Mainly expressed in the filaments of flowers, the shoot apex regions and pollen. Also present in leaves. {ECO:0000269|PubMed:20706774}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds DNA on 5'-ACTCTAC-3' and promotes auxin homeostasis-regulating gene expression (e.g. YUC genes), as well as genes affecting stamen development, cell expansion and timing of flowering. Synergistically with other SHI-related proteins, regulates gynoecium, stamen and leaf development in a dose-dependent manner, controlling apical-basal patterning. Promotes style and stigma formation, and influences vascular development during gynoecium development. May also have a role in the formation and/or maintenance of the shoot apical meristem (SAM). Regulates anther dehiscence and floral development. {ECO:0000269|PubMed:16740146, ECO:0000269|PubMed:20706774}. | |||||
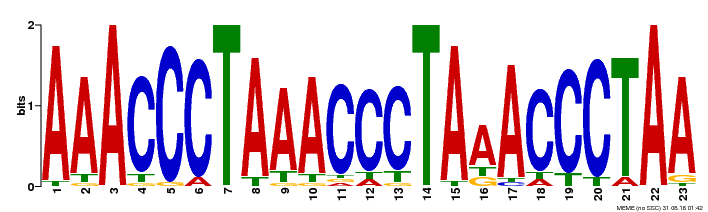
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00152 | DAP | Transfer from AT1G19790 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Medtr3g112510.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | DQ016337 | 4e-45 | DQ016337.1 Cicer reticulatum clone Crt93 microsatellite sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_013462189.1 | 0.0 | protein SHI RELATED SEQUENCE 7 | ||||
| Swissprot | Q9FXH7 | 4e-69 | SRS7_ARATH; Protein SHI RELATED SEQUENCE 7 | ||||
| TrEMBL | A0A072V2G0 | 0.0 | A0A072V2G0_MEDTR; Putative transcription factor STY-LRP1 family | ||||
| STRING | GLYMA04G03000.1 | 7e-94 | (Glycine max) | ||||
| STRING | GLYMA06G03030.1 | 6e-94 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF882 | 34 | 125 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G19790.2 | 2e-64 | SHI-related sequence 7 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Medtr3g112510.1 |
| Entrez Gene | 25490247 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



