| Signature Domain? help Back to Top |
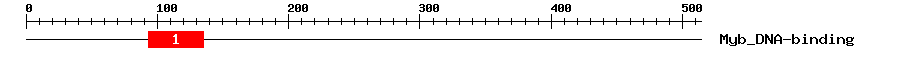 |
| No. |
Domain |
Score |
E-value |
Start |
End |
HMM Start |
HMM End |
| 1 | Myb_DNA-binding | 38.9 | 2e-12 | 93 | 135 | 3 | 46 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
W++ E++ll++a+ ++G g+W+ +a+ +g +++ qc++++++
Medtr3g081360.1 93 DWSAGEEKLLIEAIDMYGFGNWNGVAENVG-TKSKSQCIDHYNS 135
6*****************************.***********97 PP
|
| Publications
? help Back to Top |
- Vlachonasios KE,Thomashow MF,Triezenberg SJ
Disruption mutations of ADA2b and GCN5 transcriptional adaptor genes dramatically affect Arabidopsis growth, development, and gene expression.
Plant Cell, 2003. 15(3): p. 626-38
[PMID:12615937] - Sieberer T,Hauser MT,Seifert GJ,Luschnig C
PROPORZ1, a putative Arabidopsis transcriptional adaptor protein, mediates auxin and cytokinin signals in the control of cell proliferation.
Curr. Biol., 2003. 13(10): p. 837-42
[PMID:12747832] - Kikuchi S, et al.
Collection, mapping, and annotation of over 28,000 cDNA clones from japonica rice.
Science, 2003. 301(5631): p. 376-9
[PMID:12869764] - Kerschen A,Napoli CA,Jorgensen RA,Müller AE
Effectiveness of RNA interference in transgenic plants.
FEBS Lett., 2004. 566(1-3): p. 223-8
[PMID:15147899] - Hark AT, et al.
Two Arabidopsis orthologs of the transcriptional coactivator ADA2 have distinct biological functions.
Biochim. Biophys. Acta, 2009. 1789(2): p. 117-24
[PMID:18929690] - Anzola JM, et al.
Putative Arabidopsis transcriptional adaptor protein (PROPORZ1) is required to modulate histone acetylation in response to auxin.
Proc. Natl. Acad. Sci. U.S.A., 2010. 107(22): p. 10308-13
[PMID:20479223] - Kaldis A,Tsementzi D,Tanriverdi O,Vlachonasios KE
Arabidopsis thaliana transcriptional co-activators ADA2b and SGF29a are implicated in salt stress responses.
Planta, 2011. 233(4): p. 749-62
[PMID:21193996] - Vlachonasios KE,Kaldis A,Nikoloudi A,Tsementzi D
The role of transcriptional coactivator ADA2b in Arabidopsis abiotic stress responses.
Plant Signal Behav, 2011. 6(10): p. 1475-8
[PMID:21897124] - Young ND, et al.
The Medicago genome provides insight into the evolution of rhizobial symbioses.
Nature, 2011. 480(7378): p. 520-4
[PMID:22089132] - Heyndrickx KS,Vandepoele K
Systematic identification of functional plant modules through the integration of complementary data sources.
Plant Physiol., 2012. 159(3): p. 884-901
[PMID:22589469] - Kazama D,Kurusu T,Mitsuda N,Ohme-Takagi M,Tada Y
Involvement of elevated proline accumulation in enhanced osmotic stress tolerance in Arabidopsis conferred by chimeric repressor gene silencing technology.
Plant Signal Behav, 2014. 9(3): p. e28211
[PMID:24614501] - Kim JY,Oh JE,Noh YS,Noh B
Epigenetic control of juvenile-to-adult phase transition by the Arabidopsis SAGA-like complex.
Plant J., 2015. 83(3): p. 537-45
[PMID:26095998] - Kazama D, et al.
Identification of Chimeric Repressors that Confer Salt and Osmotic Stress Tolerance in Arabidopsis.
Plants (Basel), 2013. 2(4): p. 769-85
[PMID:27137403] - Simon MK,Skinner DJ,Gallagher TL,Gasser CS
Integument Development in Arabidopsis Depends on Interaction of YABBY Protein INNER NO OUTER with Coactivators and Corepressors.
Genetics, 2017. 207(4): p. 1489-1500
[PMID:28971961] - Lai J, et al.
The Transcriptional Coactivator ADA2b Recruits a Structural Maintenance Protein to Double-Strand Breaks during DNA Repair in Plants.
Plant Physiol., 2018. 176(4): p. 2613-2622
[PMID:29463775] - Kotak J, et al.
The histone acetyltransferase GCN5 and the transcriptional coactivator ADA2b affect leaf development and trichome morphogenesis in Arabidopsis.
Planta, 2018. 248(3): p. 613-628
[PMID:29846775]
|




