 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Medtr1g100970.4 | ||||||||
| Common Name | MTR_1g100970 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Trifolieae; Medicago
|
||||||||
| Family | Nin-like | ||||||||
| Protein Properties | Length: 692aa MW: 76057.1 Da PI: 6.2433 | ||||||||
| Description | Nin-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | RWP-RK | 91.1 | 8.2e-29 | 283 | 333 | 2 | 52 |
RWP-RK 2 ekeisledlskyFslpikdAAkeLgvclTvLKriCRqyGIkRWPhRkiksl 52
ek+isle+l++yF++++kdAAk+Lgvc+T++KriCRq+GI+RWP+Rki+++
Medtr1g100970.4 283 EKSISLEVLQRYFAGSLKDAAKSLGVCPTTMKRICRQHGISRWPSRKINKV 333
799**********************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51519 | 17.461 | 272 | 353 | IPR003035 | RWP-RK domain |
| Pfam | PF02042 | 1.1E-25 | 286 | 333 | IPR003035 | RWP-RK domain |
| SuperFamily | SSF54277 | 3.92E-22 | 592 | 681 | No hit | No description |
| SMART | SM00666 | 2.6E-25 | 595 | 678 | IPR000270 | PB1 domain |
| Gene3D | G3DSA:3.10.20.240 | 2.6E-26 | 595 | 677 | No hit | No description |
| Pfam | PF00564 | 7.5E-19 | 595 | 677 | IPR000270 | PB1 domain |
| PROSITE profile | PS51745 | 24.446 | 595 | 678 | IPR000270 | PB1 domain |
| CDD | cd06407 | 1.57E-33 | 596 | 677 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0010118 | Biological Process | stomatal movement | ||||
| GO:0010167 | Biological Process | response to nitrate | ||||
| GO:0042128 | Biological Process | nitrate assimilation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 692 aa Download sequence Send to blast |
MLLNQICNEG RQNALSEILE ILTVVCETHN LPLAQTWVPC RHRSVLAHGG GFKKSCSSFD 60 GSCMGQVCMS TTEAAAYIVD AHLWGFREAC VEHHLQQGQG VAGRAFLSQT MSFCTNITQF 120 CKTDYPLVHY ALMFGLTSCF AICLRSFHTG NDDYVLEFFL PPGITEFHEQ KTLLGSIFST 180 MKQHFQSLNI AAGVELEENG SIEIIEATDE GIRLRTESIP IAQSIKSPPR PDASPNMEDE 240 EEGVPQDPSE VRGENVGGSV DPMSTLGNKN IKKPSERKRG KTEKSISLEV LQRYFAGSLK 300 DAAKSLGVCP TTMKRICRQH GISRWPSRKI NKVNRSLSKL KRVIDSVQRA EGAFDLNSLG 360 NNQLPIVSSF PEPSTLNKSS QQGSLSNRPS EPQMKENEFD ASKVSETNLQ IVMENQLLGG 420 RKHSLEKEVI NGKGVTIQEI GKDRKRNRTR SGSSEDSTNP TSHGSCHGSP PIEIPTIKDL 480 FIPSNNEQHV VLRRSPEPGM QPTNALNSPT AHRMVDNVIA ELQEPFGGLL IEDAGSSKDL 540 RNLCPSVAEA ILEDMAPEAC GNNFPGSSHL APPKQCIDTI NNSATPFAAR KEMKTVTIKA 600 TYREDIIRFR VSLNCGIVEL KEEVSKRLKL EVGTFDIKYM DDDNEWVLIA CDADLQECMY 660 LSRSSGGSNI IRVLVHDITS NLGSSCESSG E* |
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, stems, leaves, flowers and siliques. Detected in root hairs, emerging secondary roots, vascular tissues, leaf parenchyma cells and stomata. {ECO:0000269|PubMed:18826430}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in regulation of nitrate assimilation and in transduction of the nitrate signal. {ECO:0000269|PubMed:18826430}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00449 | DAP | Transfer from AT4G24020 | Download |
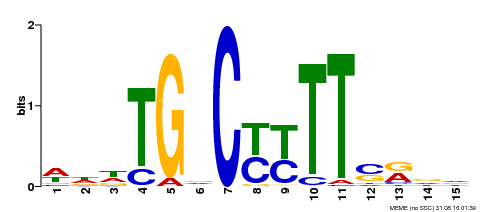
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Medtr1g100970.4 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Not regulated by the N source or by nitrate. {ECO:0000269|PubMed:18826430}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC146755 | 0.0 | AC146755.23 Medicago truncatula clone mth2-15i15, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_013469658.1 | 0.0 | protein NLP6 isoform X2 | ||||
| Swissprot | Q84TH9 | 0.0 | NLP7_ARATH; Protein NLP7 | ||||
| TrEMBL | A0A072W053 | 0.0 | A0A072W053_MEDTR; Plant regulator RWP-RK family protein | ||||
| STRING | AES62521 | 0.0 | (Medicago truncatula) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G24020.1 | 0.0 | NIN like protein 7 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Medtr1g100970.4 |
| Entrez Gene | 11421138 |



