 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Medtr1g100970.3 | ||||||||
| Common Name | MTR_1g100970 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Trifolieae; Medicago
|
||||||||
| Family | Nin-like | ||||||||
| Protein Properties | Length: 900aa MW: 99745.1 Da PI: 6.0679 | ||||||||
| Description | Nin-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | RWP-RK | 90.7 | 1.1e-28 | 491 | 541 | 2 | 52 |
RWP-RK 2 ekeisledlskyFslpikdAAkeLgvclTvLKriCRqyGIkRWPhRkiksl 52
ek+isle+l++yF++++kdAAk+Lgvc+T++KriCRq+GI+RWP+Rki+++
Medtr1g100970.3 491 EKSISLEVLQRYFAGSLKDAAKSLGVCPTTMKRICRQHGISRWPSRKINKV 541
799**********************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51519 | 17.461 | 480 | 561 | IPR003035 | RWP-RK domain |
| Pfam | PF02042 | 1.6E-25 | 494 | 541 | IPR003035 | RWP-RK domain |
| SuperFamily | SSF54277 | 5.56E-22 | 800 | 889 | No hit | No description |
| SMART | SM00666 | 2.6E-25 | 803 | 886 | IPR000270 | PB1 domain |
| Pfam | PF00564 | 1.1E-18 | 803 | 885 | IPR000270 | PB1 domain |
| Gene3D | G3DSA:3.10.20.240 | 3.7E-26 | 803 | 885 | No hit | No description |
| PROSITE profile | PS51745 | 24.446 | 803 | 886 | IPR000270 | PB1 domain |
| CDD | cd06407 | 3.65E-33 | 804 | 885 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0010118 | Biological Process | stomatal movement | ||||
| GO:0010167 | Biological Process | response to nitrate | ||||
| GO:0042128 | Biological Process | nitrate assimilation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 900 aa Download sequence Send to blast |
MASWLELPGD SNSITEKLVE KDDNKTLLPP LLPPIENLDG YCVIKEKMTQ ALRYFKEWTE 60 LNVLAQVWAP VRNGNRYVLT TSGQPFVLDP HSNGLNQYRT VSLMYMFSVD GENDGTLGLP 120 GRVFQQKLPE WSPNVLYYSN KEYPRRDHAQ HYNVRGTLAL PVFEPSLQSC IGVIELIMTS 180 LKINYAPEVE KICKALEAVN LRSSEFLDHP FTQICNEGRQ NALSEILEIL TVVCETHNLP 240 LAQTWVPCRH RSVLAHGGGF KKSCSSFDGS CMGQVCMSTT EAAAYIVDAH LWGFREACVE 300 HHLQQGQGVA GRAFLSQTMS FCTNITQFCK TDYPLVHYAL MFGLTSCFAI CLRSFHTGND 360 DYVLEFFLPP GITEFHEQKT LLGSIFSTMK QHFQSLNIAA GVELEENGSI EIIEATDEGI 420 RLRTESIPIA QSIKSPPRPD ASPNMEDEEE GVPQDPSEVR GENVGGSVDP MSTLGNKNIK 480 KPSERKRGKT EKSISLEVLQ RYFAGSLKDA AKSLGVCPTT MKRICRQHGI SRWPSRKINK 540 VNRSLSKLKR VIDSVQRAEG AFDLNSLGNN QLPIVSSFPE PSTLNKSSQQ GSLSNRPSEP 600 QMKENEFDAS KVSETNLQIV MENQLLGGRK HSLEKEVING KGVTIQEIGK DRKRNRTRSG 660 SSEDSTNPTS HGSCHGSPPI EIPTIKDLFI PSNNEQHVVL RRSPEPGMQP TNALNSPTAH 720 RMVDNVIAEL QEPFGGLLIE DAGSSKDLRN LCPSVAEAIL EDMAPEACGN NFPGSSHLAP 780 PKQCIDTINN SATPFAARKE MKTVTIKATY REDIIRFRVS LNCGIVELKE EVSKRLKLEV 840 GTFDIKYMDD DNEWVLIACD ADLQECMYLS RSSGGSNIIR VLVHDITSNL GSSCESSGE* |
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, stems, leaves, flowers and siliques. Detected in root hairs, emerging secondary roots, vascular tissues, leaf parenchyma cells and stomata. {ECO:0000269|PubMed:18826430}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in regulation of nitrate assimilation and in transduction of the nitrate signal. {ECO:0000269|PubMed:18826430}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00449 | DAP | Transfer from AT4G24020 | Download |
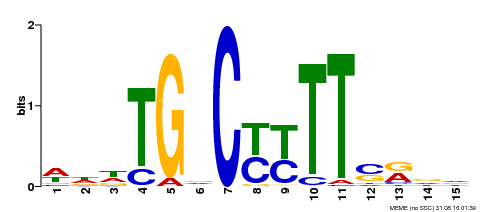
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Medtr1g100970.3 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Not regulated by the N source or by nitrate. {ECO:0000269|PubMed:18826430}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC146755 | 0.0 | AC146755.23 Medicago truncatula clone mth2-15i15, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_013469658.1 | 0.0 | protein NLP6 isoform X2 | ||||
| Swissprot | Q84TH9 | 0.0 | NLP7_ARATH; Protein NLP7 | ||||
| TrEMBL | A0A072VPV2 | 0.0 | A0A072VPV2_MEDTR; Plant regulator RWP-RK family protein | ||||
| STRING | AES62521 | 0.0 | (Medicago truncatula) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G24020.1 | 0.0 | NIN like protein 7 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Medtr1g100970.3 |
| Entrez Gene | 11421138 |



