 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Medtr1g086250.1 | ||||||||
| Common Name | MTR_1g086250 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Trifolieae; Medicago
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 1004aa MW: 111555 Da PI: 6.1171 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 127.8 | 4.2e-40 | 146 | 223 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cqve+C adls+ k+yhrrhkvCe+hska+ +lv + +qrfCqqCsrfh l+efDe+krsCrrrLa+hn+rrrk+++
Medtr1g086250.1 146 VCQVEDCGADLSRGKDYHRRHKVCEMHSKASRALVGNAMQRFCQQCSRFHILEEFDEGKRSCRRRLAGHNKRRRKTNQ 223
6**************************************************************************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 1.3E-32 | 140 | 208 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 31.778 | 144 | 221 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 2.75E-37 | 145 | 225 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 5.5E-29 | 147 | 220 | IPR004333 | Transcription factor, SBP-box |
| Gene3D | G3DSA:1.25.40.20 | 4.3E-8 | 743 | 890 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 7.6E-9 | 758 | 889 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 8.69E-8 | 777 | 888 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0016021 | Cellular Component | integral component of membrane | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1004 aa Download sequence Send to blast |
MGERLGAENY HFYGVGGSSD LSGMGKRSRE WNLNDWRWDG DLFIASRVNQ VQAESLRVGQ 60 QFFPLGSGIP VVGGSSNTSS SCSEEGDLEK GNKEGEKKRR VIVLEDDGLN DKAGALSLNL 120 AGHVSPVVER DGKKSRGAGG TSNRAVCQVE DCGADLSRGK DYHRRHKVCE MHSKASRALV 180 GNAMQRFCQQ CSRFHILEEF DEGKRSCRRR LAGHNKRRRK TNQEAVPNGS PTNDDQTSSY 240 LLISLLKILS NMHSDRSDQP TDQDLLTHLL RSLASQNDEQ GSKNLSNLLR EQENLLREGG 300 SSRNSGMVSA LFSNGSQGSP TVITQHQPVS MNQMQQEMVH THDVRTSDHQ LISSIKPSIS 360 NSPPAYSETR DSSGQTKMNN FDLNDIYVDS DDGTEDLERL PVSTNLATSS VDYPWTQQDS 420 HQSSPAQTSG NSDSASAQSP SSSSGEAQSR TDRIVFKLFG KEPNEFPLVL RAQILDWLSQ 480 SPTDIESYIR PGCIVLTIYL RQAEAVWEEL CCDLTSSLIK LLDVSDDTFW KTGWVHIRVQ 540 HQMAFIFNGQ VVIDTSLPFR SNNYSKIWTV SPIAVPASKR AQFSVKGVNL MRPATRLMCA 600 LEGKYLVCED AHESTDQYSE ELDELQCIQF SCSVPVSNGR GFIEIEDQGL SSSFFPFIVA 660 EEDVCTEIRV LEPLLESSET DPDIEGTGKI KAKSQAMDFI HEMGWLLHRS QLKYRMVNLN 720 SGVDLFPLQR FTWLMEFSMD HDWCAVVKKL LNLLLDETVN KGDHPTLYQA LSEMGLLHRA 780 VRRNSKQLVE LLLRYVPDNT SDELGPEDKA LVGGKNHSYL FRPDAVGPAG LTPLHIAAGK 840 DGSEDVLDAL TNDPCMVGIE AWKNARDSTG STPEDYARLR GHYTYIHLVQ KKINKTQGAA 900 HVVVEIPSNM TESNKNPKQN ESFTSLEIGK AEVRRSQGNC KLCDTKISCR TAVGRSMVYR 960 PAMLSMVAIA AVCVCVALLF KSSPEVLYMF RPFRWESLDF GTS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 1e-30 | 137 | 220 | 1 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Mtr.4233 | 0.0 | flower| leaf| pod| root| seed| stem | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Expressed constitutively during plant development. {ECO:0000269|PubMed:10524240}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' of AP1 promoter. Binds specifically to the 5'-GTAC-3' core sequence. {ECO:0000269|PubMed:10524240, ECO:0000269|PubMed:16095614}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00633 | PBM | Transfer from PK06791.1 | Download |
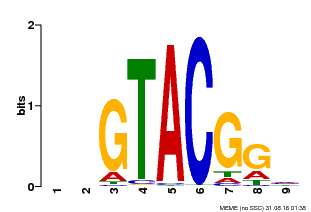
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Medtr1g086250.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003591325.1 | 0.0 | squamosa promoter-binding-like protein 1 | ||||
| Swissprot | Q9SMX9 | 0.0 | SPL1_ARATH; Squamosa promoter-binding-like protein 1 | ||||
| TrEMBL | G7IBS5 | 0.0 | G7IBS5_MEDTR; Putative transcription factor SBP family | ||||
| STRING | AES61576 | 0.0 | (Medicago truncatula) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1539 | 34 | 95 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Medtr1g086250.1 |
| Entrez Gene | 11412439 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



