 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Medtr1g017080.1 | ||||||||
| Common Name | MTR_1g017080 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Trifolieae; Medicago
|
||||||||
| Family | TALE | ||||||||
| Protein Properties | Length: 380aa MW: 43695.7 Da PI: 6.8941 | ||||||||
| Description | TALE family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 28.2 | 3.1e-09 | 306 | 341 | 20 | 55 |
HHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHH CS
Homeobox 20 eknrypsaeereeLAkklgLterqVkvWFqNrRake 55
+k +yps++e+ LA+++gL+++q+ +WF N+R ++
Medtr1g017080.1 306 YKWPYPSESEKVALAESTGLDQKQINNWFINQRKRH 341
4679*****************************885 PP
| |||||||
| 2 | ELK | 36 | 1.5e-12 | 261 | 282 | 1 | 22 |
ELK 1 ELKhqLlrKYsgyLgsLkqEFs 22
ELK++Ll+KYsgyL+sLkqE+s
Medtr1g017080.1 261 ELKNHLLKKYSGYLSSLKQELS 282
9*******************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01255 | 8.6E-21 | 117 | 161 | IPR005540 | KNOX1 |
| Pfam | PF03790 | 4.7E-22 | 118 | 160 | IPR005540 | KNOX1 |
| SMART | SM01256 | 3.7E-30 | 168 | 219 | IPR005541 | KNOX2 |
| Pfam | PF03791 | 6.5E-25 | 172 | 218 | IPR005541 | KNOX2 |
| SMART | SM01188 | 1.0E-6 | 261 | 282 | IPR005539 | ELK domain |
| Pfam | PF03789 | 7.5E-10 | 261 | 282 | IPR005539 | ELK domain |
| PROSITE profile | PS51213 | 11.243 | 261 | 281 | IPR005539 | ELK domain |
| PROSITE profile | PS50071 | 13.119 | 281 | 344 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 6.42E-20 | 282 | 356 | IPR009057 | Homeodomain-like |
| SMART | SM00389 | 1.8E-13 | 283 | 348 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 1.3E-27 | 286 | 347 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 7.38E-12 | 293 | 345 | No hit | No description |
| Pfam | PF05920 | 1.1E-16 | 301 | 340 | IPR008422 | Homeobox KN domain |
| PROSITE pattern | PS00027 | 0 | 319 | 342 | IPR017970 | Homeobox, conserved site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0001708 | Biological Process | cell fate specification | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010051 | Biological Process | xylem and phloem pattern formation | ||||
| GO:0010089 | Biological Process | xylem development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 380 aa Download sequence Send to blast |
MEEYTNNPNP NPNSRPNFLY SIASGNNQHQ HQHNHQHNQI PMNNFHGSDN CFQSDQVQHQ 60 QHSAVKTEAN STSQLHTPIF HYPALMRTNI IPHTNIMHNH HHQGGGGSPS SSNVEAEAIK 120 AKIIAHPQYS SLLQAYMDCQ KIGAPPEVVA RLVASRQEFE ARQRSSVNSR ETSKDPELDQ 180 FMEAYYDMLV KYREELTRPI QEAMDFMRRI ETQLNTLCNG PLRIFPDDKN EGVGSSEEDQ 240 ENSGGETDQL PEIDPRAEDR ELKNHLLKKY SGYLSSLKQE LSKKKKKGKL PKEARQKLLN 300 WWELHYKWPY PSESEKVALA ESTGLDQKQI NNWFINQRKR HWKPSEDMQF MVMDGLHPQS 360 AALYMDGHYM ADGPYRLGP* |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Mtr.7182 | 0.0 | leaf| root| seed| stem | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in the apical meristems, in the newly emerged lateral primordia in the floral bud, in their vascular bundles and in the cortex parenchyma of the floral pedicle. Also present in the lateral tips of leaf primordia. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed only in the stems. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Appears to be involved in meristem formation and in the regulation of leaf morphology. Misexpression makes the leaf more compound which is always associated with growth retardation and loss of apical dominance, resulting in dwarfed, bushy plants. Probably binds to the DNA sequence 5'-TGAC-3'. | |||||
| UniProt | Probably binds to the DNA sequence 5'-TGAC-3'. | |||||
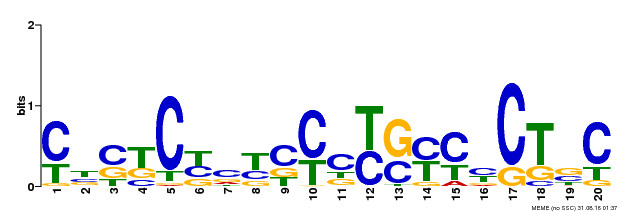
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00675 | ChIP-seq | Transfer from GRMZM2G017087 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Medtr1g017080.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | EF128057 | 0.0 | EF128057.1 Medicago truncatula class I KNOX homeobox transcription factor (KNOX2) mRNA, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_013465932.1 | 0.0 | homeotic protein knotted-1 | ||||
| Swissprot | O04135 | 1e-163 | KNAP2_MALDO; Homeobox protein knotted-1-like 2 | ||||
| Swissprot | Q41330 | 1e-164 | KN1_SOLLC; Homeotic protein knotted-1 | ||||
| TrEMBL | X2D315 | 0.0 | X2D315_MEDTR; BP | ||||
| STRING | XP_004498934.1 | 0.0 | (Cicer arietinum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF4584 | 32 | 46 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G08150.1 | 1e-132 | KNOTTED-like from Arabidopsis thaliana | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Medtr1g017080.1 |
| Entrez Gene | 25482021 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



