 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 106146 | ||||||||
| Common Name | MICPUN_106146 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; prasinophytes; Mamiellophyceae; Mamiellales; Mamiellaceae; Micromonas
|
||||||||
| Family | ARR-B | ||||||||
| Protein Properties | Length: 489aa MW: 50680.9 Da PI: 6.863 | ||||||||
| Description | ARR-B family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 91.2 | 8.9e-29 | 224 | 277 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kpr++W+ eLH++Fv av+qL G++kA+Pk+il+lm+v+gLt+e+v+SHLQkYRl
106146 224 KPRVVWSAELHQQFVTAVNQL-GIDKAVPKRILDLMGVQGLTRENVASHLQKYRL 277
79*******************.********************************8 PP
| |||||||
| 2 | Response_reg | 88.1 | 2.5e-29 | 16 | 124 | 1 | 109 |
EEEESSSHHHHHHHHHHHHHTTCEEEEEESSHHHHHHHHHHHH..ESEEEEESSCTTSEHHHHHHHHHHHTTTSEEEEEESTTTHHHHHHHHHTTES CS
Response_reg 1 vlivdDeplvrellrqalekegyeevaeaddgeealellkekd..pDlillDiempgmdGlellkeireeepklpiivvtahgeeedalealkaGak 95
vl+vdD+pl +++++++l++ +y ev++++ g eal++l+e++ +D++l D+ mp+mdG++ll++i+ e ++p+++++a+ + + +l+ + Ga
106146 16 VLVVDDDPLCLKIVEKMLKRCQY-EVTTFSRGAEALKTLRERKddFDIVLSDVHMPDMDGFKLLEHIALEL-DIPVMMMSANCATDVVLRGIIHGAV 110
89*********************.***************777777**********************8765.8************************ PP
EEEESS--HHHHHH CS
Response_reg 96 dflsKpfdpeelvk 109
d+l Kp+ +eel++
106146 111 DYLLKPVRIEELRN 124
************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:3.40.50.2300 | 2.6E-42 | 13 | 138 | No hit | No description |
| SuperFamily | SSF52172 | 1.57E-37 | 13 | 137 | IPR011006 | CheY-like superfamily |
| SMART | SM00448 | 6.9E-33 | 14 | 126 | IPR001789 | Signal transduction response regulator, receiver domain |
| PROSITE profile | PS50110 | 42.673 | 15 | 130 | IPR001789 | Signal transduction response regulator, receiver domain |
| Pfam | PF00072 | 4.4E-26 | 16 | 125 | IPR001789 | Signal transduction response regulator, receiver domain |
| CDD | cd00156 | 2.05E-31 | 17 | 129 | No hit | No description |
| PROSITE profile | PS51294 | 11.312 | 221 | 280 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 6.45E-20 | 222 | 281 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 3.5E-30 | 223 | 282 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 6.9E-25 | 224 | 277 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 5.3E-8 | 226 | 276 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009873 | Biological Process | ethylene-activated signaling pathway | ||||
| GO:0010082 | Biological Process | regulation of root meristem growth | ||||
| GO:0010119 | Biological Process | regulation of stomatal movement | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0071368 | Biological Process | cellular response to cytokinin stimulus | ||||
| GO:0080113 | Biological Process | regulation of seed growth | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 489 aa Download sequence Send to blast |
MSTPAVSKGF PIGLRVLVVD DDPLCLKIVE KMLKRCQYEV TTFSRGAEAL KTLRERKDDF 60 DIVLSDVHMP DMDGFKLLEH IALELDIPVM MMSANCATDV VLRGIIHGAV DYLLKPVRIE 120 ELRNIWQHVV RRKRESSQGN LRSGEGGSNG RTVSGGSTGE GGGKDSKGSS EQHGDAKDKT 180 GSAGGSGGSS KRKKGSGKKG DEGTDEVKDG SGGDENEDSS ALKKPRVVWS AELHQQFVTA 240 VNQLGIDKAV PKRILDLMGV QGLTRENVAS HLQKYRLYLK RLQGVNSGGA PGGGPGFMSP 300 IALDGSMVQG GPGGRVGSPA IGGPNGPIMV GHGHIDPAML AGGAPQTIQM GMVYGGPGMG 360 PPQMMAPNGK GGGGMPGGYV MQPGQMMAPN GQMMPVGQMG PGGMMVQGPG GGMMQMHDGG 420 MMNGNGSYGS LQNMKQGNGV VMMPNGGMGG VDGAIPNMAT GLINGQGLPD DDVLDMFLKD 480 GLPEGEGF* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 2e-22 | 220 | 282 | 1 | 63 | ARR10-B |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specifically to the DNA sequence 5'-[AG]GATT-3'. Functions as a response regulator involved in His-to-Asp phosphorelay signal transduction system. Phosphorylation of the Asp residue in the receiver domain activates the ability of the protein to promote the transcription of target genes. Could directly activate some type-A response regulators in response to cytokinins. Involved in the expression of nuclear genes for components of mitochondrial complex I. Promotes cytokinin-mediated leaf longevity. Involved in the ethylene signaling pathway in an ETR1-dependent manner and in the cytokinin signaling pathway. {ECO:0000269|PubMed:11370868, ECO:0000269|PubMed:11574878, ECO:0000269|PubMed:15282545, ECO:0000269|PubMed:16407152}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00054 | PBM | Transfer from AT4G16110 | Download |
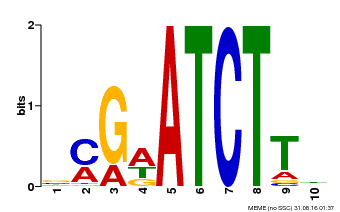
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | CP001576 | 0.0 | CP001576.1 Micromonas sp. RCC299 chromosome 10, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002508755.1 | 0.0 | g2-like myb-transcription factor with a n-terminal response regulator receiver domain | ||||
| Swissprot | Q9ZWJ9 | 1e-101 | ARR2_ARATH; Two-component response regulator ARR2 | ||||
| TrEMBL | C1FH07 | 0.0 | C1FH07_MICCC; G2-like myb-transcription factor with a n-terminal response regulator receiver domain | ||||
| STRING | XP_002508755.1 | 0.0 | (Micromonas sp. RCC299) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Chlorophytae | OGCP1220 | 15 | 16 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G16110.1 | 7e-94 | response regulator 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | 106146 |
| Entrez Gene | 8247032 |



