 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 50595 | ||||||||
| Common Name | MICPUCDRAFT_50595 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; prasinophytes; Mamiellophyceae; Mamiellales; Mamiellaceae; Micromonas; Micromonas pusilla
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1151aa MW: 116379 Da PI: 6.3611 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 96.9 | 1.7e-30 | 53 | 128 | 2 | 75 |
CG-1 2 lkekkrwlkneeiaaiLenfekheltl..elktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKv 75
+++++rwlkn+e+++iL n++ ++++l +++ +p+ gsl+L++rk+vr+frkDG++w+kkkdgktvrE+hekLK+
50595 53 QQSQTRWLKNTEVCDILLNHRAYDFVLspNAPIQPSAGSLFLFDRKVVRFFRKDGHEWQKKKDGKTVRETHEKLKM 128
4459*******************9998778899******************************************6 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00248 | 3600 | 41 | 79 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS51437 | 39.611 | 48 | 180 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 4.8E-28 | 51 | 149 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 3.1E-26 | 55 | 128 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:1.25.40.20 | 5.1E-15 | 62 | 74 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF81296 | 7.52E-15 | 738 | 824 | IPR014756 | Immunoglobulin E-set |
| Gene3D | G3DSA:1.25.40.20 | 5.1E-15 | 1033 | 1131 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 3.26E-14 | 1036 | 1131 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 3.50E-12 | 1038 | 1129 | No hit | No description |
| PROSITE profile | PS50297 | 16.308 | 1040 | 1129 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 1.2E-4 | 1073 | 1102 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 11.381 | 1073 | 1105 | IPR002110 | Ankyrin repeat |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0050826 | Biological Process | response to freezing | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1151 aa Download sequence Send to blast |
MSGSGSGSGG RTSTLMDEGG GGDGYSPAAI VLTPPSPMGA DGASRAHIIA LLQQSQTRWL 60 KNTEVCDILL NHRAYDFVLS PNAPIQPSAG SLFLFDRKVV RFFRKDGHEW QKKKDGKTVR 120 ETHEKLKMLL RPSRVGRDGV VLVHYLKVTP GMKTERGGHG GDLSNWPAGA YPGHGFQNLF 180 GGGDSGGTTD SGLYGTLSGT FSGTISGGGT SGGAFGSRGG YSDIEIDESD LMIRSAGSSP 240 TVSQMRGAGG FGGRVGHAYA SHQGGTSVYA SSPAAAAARA AAQMQMHDAR LGRPITNDPP 300 ANVDRWWSGR GVGTGDDDEY GVNDPELEEL LRDSDMELGT PHGGAGAPGG GQGRPNELHQ 360 YLMNEIGYEG GGNAPPPPPP GSDNMRQKSW DSGLSLSVSG LTGLPSAPGE DSVHSGTIAG 420 SLGVGAPGHG QAPPPHPLAA AAAEAAAAAG PDITTSFLSP TPAPSSSADD LSGRGADRSS 480 AMRGVTSLTQ IEGKIADLQQ TLLRVASNAV HTNVEQVARL EEDLIALERG AEAMMPRAIG 540 SGSGTGGGSG GGGPGRGSRK LFGRGGDGAE HHRRGRSFDG GGGGGHRAVA QRKPPPMRWD 600 ADDDGGDSGG ADEEDDDDDD DAEKSGDDGE DPHEGLLSPL KSAAAATRRP GGAANRRSYP 660 PRHAAPPVPI ASTARGVSQS GSEMDDGATD ADDRAPSPGM AAAAMRASSA QAAQFAQRSA 720 RARAPIVTVP PTSSILWEIH DFSPEWDVES GGAKVIISGA ARPGLPEGLH LCCVFGEIEV 780 PAEQISPGVL RCRAPPRSAG RVPLYISCLG GGKRPASDIR TFEYKETSGG GAKDRRTAEV 840 RLTTGVTERD FQLRLVHLLI GADRSSVRDR SSAGGGGGGG GGGGSGGGGG GGGSGSGGGN 900 DSAAGGGGGN GGSGGGSGSG GSSNDGGGAP GASANITPAP ANPSPNPNPG ASFARKSVAA 960 LASPGTSLAD PLASLFSASP GAADDLTDED VSRVFKTALE ARLRHAISAE AKRHRVVTTG 1020 VVPNPGYVLP RSAYHRIDSG GMGLIHCVAA LGMSWAIPAM VRTGCEVNQP DRRARTALHW 1080 AAAKGHEDTV ACLLAEGANI RATARWGAGG YTAADLAAAL GHGGIAAYIS EVRSTHWFPY 1140 DPVAVVNADP * |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00501 | DAP | Transfer from AT5G09410 | Download |
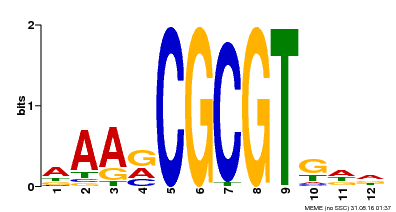
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003055680.1 | 0.0 | camta-like transcriptional regulator | ||||
| TrEMBL | C1MIJ9 | 0.0 | C1MIJ9_MICPC; Camta-like transcriptional regulator | ||||
| STRING | XP_003055680.1 | 0.0 | (Micromonas pusilla) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Chlorophytae | OGCP5132 | 8 | 9 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G09410.1 | 2e-25 | ethylene induced calmodulin binding protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | 50595 |
| Entrez Gene | 9680865 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



