 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Mapoly1089s0002.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Marchantiophyta; Marchantiopsida; Marchantiidae; Marchantiales; Marchantiaceae; Marchantia
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 597aa MW: 64604.9 Da PI: 5.4256 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 54.6 | 2.5e-17 | 18 | 65 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT Ed +lv +v+++G g+W+++ ++ g+ R++k+c++rw ++l
Mapoly1089s0002.1.p 18 KGPWTSAEDAILVAYVTKHGEGNWNSVQKHSGLYRCGKSCRLRWANHL 65
79******************************************9986 PP
| |||||||
| 2 | Myb_DNA-binding | 50.1 | 6.4e-16 | 71 | 114 | 1 | 46 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
+g++T+eE+ +++++++lG++ W++ a+ ++ gRt++++k++w++
Mapoly1089s0002.1.p 71 KGAFTPEEERMIIELHAKLGNK-WARMAAQLP-GRTDNEIKNYWNT 114
799*******************.*********.***********96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 14.794 | 15 | 65 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 3.41E-30 | 16 | 112 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 2.8E-14 | 17 | 67 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.6E-14 | 18 | 65 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.2E-23 | 19 | 72 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 9.67E-11 | 20 | 65 | No hit | No description |
| PROSITE profile | PS51294 | 25.733 | 66 | 120 | IPR017930 | Myb domain |
| SMART | SM00717 | 3.7E-16 | 70 | 118 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 5.4E-15 | 71 | 114 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 9.98E-12 | 73 | 114 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 3.4E-26 | 73 | 120 | IPR009057 | Homeodomain-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009789 | Biological Process | positive regulation of abscisic acid-activated signaling pathway | ||||
| GO:0043068 | Biological Process | positive regulation of programmed cell death | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0045926 | Biological Process | negative regulation of growth | ||||
| GO:0048235 | Biological Process | pollen sperm cell differentiation | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 597 aa Download sequence Send to blast |
MELGSAPDFS EDVGALKKGP WTSAEDAILV AYVTKHGEGN WNSVQKHSGL YRCGKSCRLR 60 WANHLRPNLK KGAFTPEEER MIIELHAKLG NKWARMAAQL PGRTDNEIKN YWNTRIKRRM 120 RAGLPVYPPE MQNPAANSQY LFEHGEMSMS SGGESECDPG SSPSTGDFQN LSVSGMHGGC 180 RMKSCSSLTG MSDLPLSSVV TQTLSAGQSI SSPGRRMKRM HRDTQCSSMS GASGGGGLFP 240 QLSDESSKTM PYFKARRSCG TRNSMRLAQI AGFPYDPDPE GLHFEGMPNH GYLNLPPFSC 300 SRPNSSLKLE LPSSQSAESA DSAGTPGSAM TSPYTAFSLP QNHHLLSSEA DSFGSNNSNN 360 SSFLQALLQE AQNLDPGRDQ IRSAELSDQL LVLTSANPPM DVSALMSPRK SRWGEDSDPT 420 TPLEGRTYSM FSEDTSPNCT SNWDETSTLQ SPLTTVSSNL QAAHVGGLKI ENSAQHDMPC 480 GNYGDEENLI SSLLDFARPD ATPVVEWYNP PDVYTLGGQP CHSLPEAIEA AFHHQDVVAE 540 LEHLGAAGHP VANHVWELGS CPWNNMPGVC QLGDLPTDCR PLTSIADQMN DPCAIC* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 6e-32 | 16 | 120 | 5 | 108 | B-MYB |
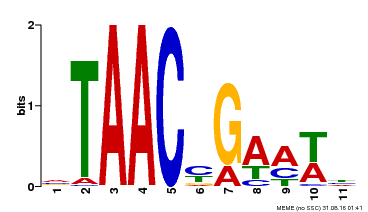
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00490 | DAP | Transfer from AT5G06100 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| TrEMBL | A0A1B4Z1R7 | 0.0 | A0A1B4Z1R7_MARPO; Myb domain protein 33-like protein | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP5 | 17 | 1784 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G06100.1 | 2e-69 | myb domain protein 33 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Mapoly1089s0002.1.p |



