 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Mapoly0001s0298.4.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Marchantiophyta; Marchantiopsida; Marchantiidae; Marchantiales; Marchantiaceae; Marchantia
|
||||||||
| Family | TCP | ||||||||
| Protein Properties | Length: 1146aa MW: 122880 Da PI: 8.0822 | ||||||||
| Description | TCP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCP | 117.5 | 1.7e-36 | 588 | 787 | 2 | 128 |
TCP 2 agkkdrhskihTkvggRdRRvRlsaecaarfFdLqdeLGfdkdsktieWLlqqakpaikeltgt.......................... 65
g+kdrhsk+ T++g+RdRRvRls+++a +f+d+qd+LGfd++sk++eWL+ +ak ai+el +
Mapoly0001s0298.4.p 588 SGGKDRHSKVNTAKGPRDRRVRLSVPTAVQFYDVQDRLGFDQPSKAVEWLIHKAKDAIDELGRIpvdgkdgprvpsvctsditrhvsiss 677
689************************************************************999999998888888888888776666 PP
TCP 66 ...............ssssaseceaesssssasnsssg.................................................kaa 91
s c+++s + + +
Mapoly0001s0298.4.p 678 sygagavsglvsgtsS------CSPAS----------SfqgpssmsygdqlsflmnpsiaqgmfaplqpnqgnfagptfgvgrmdltRPN 751
6666555555444331......32222..........022222233345555666777778999999999999***********986555 PP
TCP 92 ksaakskksqksaasalnlak.esrakarararertre 128
+ +a++ +s++sa + la+ esr kar+rarert++
Mapoly0001s0298.4.p 752 SGSAEGGRSNESAG--SPLARvESRTKARERARERTKK 787
55556666666666..899999**************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03634 | 7.8E-34 | 589 | 787 | IPR005333 | Transcription factor, TCP |
| PROSITE profile | PS51369 | 32.971 | 590 | 648 | IPR017887 | Transcription factor TCP subgroup |
| PROSITE profile | PS51370 | 9.434 | 771 | 788 | IPR017888 | CYC/TB1, R domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0030154 | Biological Process | cell differentiation | ||||
| GO:0045962 | Biological Process | positive regulation of development, heterochronic | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1146 aa Download sequence Send to blast |
MLAWARSVDS STSCHARSTS WRIGFEGNCW PVERDLQDVC NRQCSDSTGY LEVSVRAAKQ 60 PQSELCFSVA NGSKFESKVD ERARDGERFR IERDQIRQYA QSIYRTLADP QAAAVPTEQR 120 SSGLRSSIPG GQLACDVSKY RVQTASANQL HPNSNDSFES INFSPSSPIT DHQRELRRTE 180 IEAGLEPEGT RRQNVGEEAS DQINRATSLK NGFPLAERAS APEKFIAEGL DDSEPARHGR 240 GGPQNKRSSI WAKNVELVGH GPFAFRENIV ECSTHIGVSE KEGFVLHNVI DAQASHGRAK 300 EPTADSGTPR SQNSHQRTIH PQGRQDRAKT ANAIPSSPQS QALTDRVSRG AASDALGTDL 360 LPQSIGSDLL QQGTTPARVE RVQNISRTRS GGERDVEDTL KGRRYPERIG DLRATTVATA 420 GRQHKAERSG EIPPVRTSQE TVVGTAGVLT IPERIGGTAE DLTAREIVDV SESHSVSERV 480 GGRAETHSAQ DGGSAEGFTI RERTGGSPSE GNKIPERIGA VSDRDSAGKV QEKGKEEKLH 540 PREEGHQRQS ESSDERRLRA RIDGSGGSAS GGGDEIEEPK VVRPARPSGG KDRHSKVNTA 600 KGPRDRRVRL SVPTAVQFYD VQDRLGFDQP SKAVEWLIHK AKDAIDELGR IPVDGKDGPR 660 VPSVCTSDIT RHVSISSSYG AGAVSGLVSG TSSCSPASSF QGPSSMSYGD QLSFLMNPSI 720 AQGMFAPLQP NQGNFAGPTF GVGRMDLTRP NSGSAEGGRS NESAGSPLAR VESRTKARER 780 ARERTKKGPG RDGNQGKVTS GGSPSSSQSP PFHPNPAFNS FNASNSIQSS GHGLSLQSPH 840 QSALLPAAYD GSPFSRIMQG SHQATFQNMI MHNPPLDCQF GDQTQSPLHN AFLVQPQASL 900 PSPSHTMNTL QHPHLQTMQV NAQGALHLGT NFNPGPCSSS FAEASMSSPH YPPVPTLPIS 960 QSLSIPQLSS LGMSPASSLG PHYPTGSPTP IPPGFSPGPP ASAFRAHSLS ISQCSNPNQN 1020 HQQQQQQLQG LSRVQRPVPS FSTTPSTSHA GSSSSRAHAH PYYSGNSSTA LMGCLQSMPS 1080 FAPRSNVGYE HGQRVRPMMS PNRIHSYQAS NSQIPARIHG LDDMEELEEE PNPATPSSPH 1140 LSPHP* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5zkt_A | 7e-20 | 595 | 648 | 1 | 54 | Putative transcription factor PCF6 |
| 5zkt_B | 7e-20 | 595 | 648 | 1 | 54 | Putative transcription factor PCF6 |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
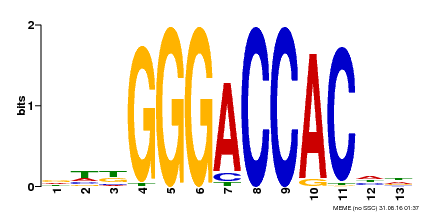
| Motif ID | Method | Source | Motif file |
| MP00203 | DAP | Transfer from AT1G53230 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| TrEMBL | A0A176W872 | 0.0 | A0A176W872_MARPO; Uncharacterized protein | ||||
| TrEMBL | A0A2R6XW17 | 0.0 | A0A2R6XW17_MARPO; Uncharacterized protein | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G18390.2 | 6e-30 | TEOSINTE BRANCHED 1, cycloidea and PCF transcription factor 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Mapoly0001s0298.4.p |



