 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Mapoly0001s0298.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Marchantiophyta; Marchantiopsida; Marchantiidae; Marchantiales; Marchantiaceae; Marchantia
|
||||||||
| Family | TCP | ||||||||
| Protein Properties | Length: 1149aa MW: 123362 Da PI: 8.3328 | ||||||||
| Description | TCP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCP | 117.5 | 1.7e-36 | 591 | 790 | 2 | 128 |
TCP 2 agkkdrhskihTkvggRdRRvRlsaecaarfFdLqdeLGfdkdsktieWLlqqakpaikeltgt.......................... 65
g+kdrhsk+ T++g+RdRRvRls+++a +f+d+qd+LGfd++sk++eWL+ +ak ai+el +
Mapoly0001s0298.1.p 591 SGGKDRHSKVNTAKGPRDRRVRLSVPTAVQFYDVQDRLGFDQPSKAVEWLIHKAKDAIDELGRIpvdgkdgprvpsvctsditrhvsiss 680
689************************************************************999999998888888888888776666 PP
TCP 66 ...............ssssaseceaesssssasnsssg.................................................kaa 91
s c+++s + + +
Mapoly0001s0298.1.p 681 sygagavsglvsgtsS------CSPAS----------SfqgpssmsygdqlsflmnpsiaqgmfaplqpnqgnfagptfgvgrmdltRPN 754
6666555555444331......32222..........022222233345555666777778999999999999***********986555 PP
TCP 92 ksaakskksqksaasalnlak.esrakarararertre 128
+ +a++ +s++sa + la+ esr kar+rarert++
Mapoly0001s0298.1.p 755 SGSAEGGRSNESAG--SPLARvESRTKARERARERTKK 790
55556666666666..899999**************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03634 | 7.8E-34 | 592 | 790 | IPR005333 | Transcription factor, TCP |
| PROSITE profile | PS51369 | 32.971 | 593 | 651 | IPR017887 | Transcription factor TCP subgroup |
| PROSITE profile | PS51370 | 9.434 | 774 | 791 | IPR017888 | CYC/TB1, R domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0030154 | Biological Process | cell differentiation | ||||
| GO:0045962 | Biological Process | positive regulation of development, heterochronic | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1149 aa Download sequence Send to blast |
MILSRLREGV QNSGWTSRLR STSWRIGFEG NCWPVERDLQ DVCNRQCSDS TGYLEVSVRA 60 AKQPQSELCF SVANGSKFES KVDERARDGE RFRIERDQIR QYAQSIYRTL ADPQAAAVPT 120 EQRSSGLRSS IPGGQLACDV SKYRVQTASA NQLHPNSNDS FESINFSPSS PITDHQRELR 180 RTEIEAGLEP EGTRRQNVGE EASDQINRAT SLKNGFPLAE RASAPEKFIA EGLDDSEPAR 240 HGRGGPQNKR SSIWAKNVEL VGHGPFAFRE NIVECSTHIG VSEKEGFVLH NVIDAQASHG 300 RAKEPTADSG TPRSQNSHQR TIHPQGRQDR AKTANAIPSS PQSQALTDRV SRGAASDALG 360 TDLLPQSIGS DLLQQGTTPA RVERVQNISR TRSGGERDVE DTLKGRRYPE RIGDLRATTV 420 ATAGRQHKAE RSGEIPPVRT SQETVVGTAG VLTIPERIGG TAEDLTAREI VDVSESHSVS 480 ERVGGRAETH SAQDGGSAEG FTIRERTGGS PSEGNKIPER IGAVSDRDSA GKVQEKGKEE 540 KLHPREEGHQ RQSESSDERR LRARIDGSGG SASGGGDEIE EPKVVRPARP SGGKDRHSKV 600 NTAKGPRDRR VRLSVPTAVQ FYDVQDRLGF DQPSKAVEWL IHKAKDAIDE LGRIPVDGKD 660 GPRVPSVCTS DITRHVSISS SYGAGAVSGL VSGTSSCSPA SSFQGPSSMS YGDQLSFLMN 720 PSIAQGMFAP LQPNQGNFAG PTFGVGRMDL TRPNSGSAEG GRSNESAGSP LARVESRTKA 780 RERARERTKK GPGRDGNQGK VTSGGSPSSS QSPPFHPNPA FNSFNASNSI QSSGHGLSLQ 840 SPHQSALLPA AYDGSPFSRI MQGSHQATFQ NMIMHNPPLD CQFGDQTQSP LHNAFLVQPQ 900 ASLPSPSHTM NTLQHPHLQT MQVNAQGALH LGTNFNPGPC SSSFAEASMS SPHYPPVPTL 960 PISQSLSIPQ LSSLGMSPAS SLGPHYPTGS PTPIPPGFSP GPPASAFRAH SLSISQCSNP 1020 NQNHQQQQQQ LQGLSRVQRP VPSFSTTPST SHAGSSSSRA HAHPYYSGNS STALMGCLQS 1080 MPSFAPRSNV GYEHGQRVRP MMSPNRIHSY QASNSQIPAR IHGLDDMEEL EEEPNPATPS 1140 SPHLSPHP* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5zkt_A | 8e-20 | 598 | 651 | 1 | 54 | Putative transcription factor PCF6 |
| 5zkt_B | 8e-20 | 598 | 651 | 1 | 54 | Putative transcription factor PCF6 |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
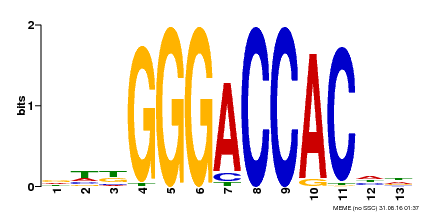
| Motif ID | Method | Source | Motif file |
| MP00203 | DAP | Transfer from AT1G53230 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| TrEMBL | A0A2R6XW12 | 0.0 | A0A2R6XW12_MARPO; Uncharacterized protein | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP180 | 15 | 163 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G18390.2 | 7e-30 | TEOSINTE BRANCHED 1, cycloidea and PCF transcription factor 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Mapoly0001s0298.1.p |



