 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010113448.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Moraceae; Morus
|
||||||||
| Family | AP2 | ||||||||
| Protein Properties | Length: 562aa MW: 61538 Da PI: 5.82 | ||||||||
| Description | AP2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 50.3 | 5.8e-16 | 378 | 429 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s y+GV++++++grW A+I + +k +lg+f t+eeAa+a++ a+ k++g
XP_010113448.1 378 SIYRGVTRHHQQGRWQARIGRVAG---NKDLYLGTFATEEEAAEAYDIAAIKFRG 429
57****************988532...5*************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00018 | 1.08E-13 | 236 | 345 | No hit | No description |
| SMART | SM00380 | 1.1E-18 | 237 | 349 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 15.158 | 237 | 343 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 1.0E-10 | 237 | 259 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 6.8E-6 | 238 | 249 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 1.0E-10 | 309 | 344 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 4.77E-8 | 313 | 345 | IPR016177 | DNA-binding domain |
| PRINTS | PR00367 | 6.8E-6 | 325 | 345 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 6.08E-18 | 378 | 439 | IPR016177 | DNA-binding domain |
| CDD | cd00018 | 5.15E-12 | 378 | 439 | No hit | No description |
| Pfam | PF00847 | 3.0E-10 | 378 | 429 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 2.7E-31 | 379 | 443 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 19.164 | 379 | 437 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 9.6E-19 | 379 | 438 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010080 | Biological Process | regulation of floral meristem growth | ||||
| GO:0010492 | Biological Process | maintenance of shoot apical meristem identity | ||||
| GO:0035265 | Biological Process | organ growth | ||||
| GO:0048364 | Biological Process | root development | ||||
| GO:0060772 | Biological Process | leaf phyllotactic patterning | ||||
| GO:0060774 | Biological Process | auxin mediated signaling pathway involved in phyllotactic patterning | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 562 aa Download sequence Send to blast |
MAPATNCNWL SFSLSPLEML RSSESTQFMA AAYENSASTA TPQYLIDNFY ANGWANPKSQ 60 VFLTEEDNQG KESQPLSNIA DDSPILTSFI DPQSQSQAVP KLEDFLGDSS SMVRYSDSQT 120 ETQDSSLTQI YDASYFADQQ QDLKVKAVTG FQAFSTNSGS EVDDSASIPR TQLACAEFAG 180 HSIESSAGNE LAFSSCPTGS LSLGVTTQSL STEKTFVPVD SDCSKKVADT FGQRTSIYRG 240 VTRHRWTGRY EAHLWDNSCR REGQARKGRQ GFDATHTPKP QMFADINGLL SNASDFLSLP 300 LTFAFAFIRQ RKSGGYDKEE KAARAYDLAA LKYWGANATT NFPVSNYSKE LEEMKHVTKQ 360 EFIASLRRKS SGFSRGASIY RGVTRHHQQG RWQARIGRVA GNKDLYLGTF ATEEEAAEAY 420 DIAAIKFRGA NAVTNFEMSR YDVEAIAKSA LPVGGAAKRL KLSLDSEDQK PSLTQDQQTQ 480 YGNSIVTTTA NNINFSTIQP ASAVPCGIPF DAALYHHNLF QHFQPNYIGG ADAAGSSPAV 540 AAPVTALISQ PAEFFVWPHQ SY |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probably acts as a transcriptional activator. Binds to the GCC-box pathogenesis-related promoter element. May be involved in the regulation of gene expression by stress factors and by components of stress signal transduction pathways (By similarity). {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00504 | DAP | Transfer from AT5G10510 | Download |
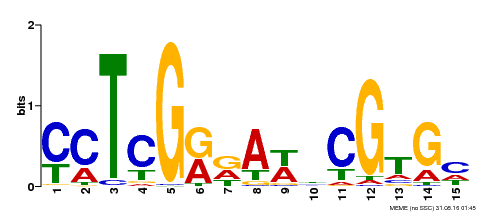
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_010113448.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024032930.1 | 0.0 | AP2-like ethylene-responsive transcription factor AIL6 | ||||
| Swissprot | Q52QU2 | 0.0 | AIL6_ARATH; AP2-like ethylene-responsive transcription factor AIL6 | ||||
| TrEMBL | W9T116 | 0.0 | W9T116_9ROSA; AP2-like ethylene-responsive transcription factor AIL6 | ||||
| STRING | XP_010113448.1 | 0.0 | (Morus notabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF4810 | 31 | 49 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G10510.3 | 0.0 | AINTEGUMENTA-like 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Entrez Gene | 21409425 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



