 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010101901.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Moraceae; Morus
|
||||||||
| Family | ARR-B | ||||||||
| Protein Properties | Length: 654aa MW: 71676.4 Da PI: 5.9565 | ||||||||
| Description | ARR-B family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 91.4 | 7.8e-29 | 204 | 257 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kpr++W+ eLH++Fv av+qL G++kA+Pk+ilelm+v+gLt+e+v+SHLQkYRl
XP_010101901.1 204 KPRVVWSVELHQQFVAAVNQL-GIDKAVPKKILELMNVPGLTRENVASHLQKYRL 257
79*******************.********************************8 PP
| |||||||
| 2 | Response_reg | 75.8 | 1.6e-25 | 26 | 134 | 1 | 109 |
EEEESSSHHHHHHHHHHHHHTTCEEEEEESSHHHHHHHHHHHH..ESEEEEESSCTTSEHHHHHHHHHHHTTTSEEEEEESTTTHHHHHHHHHTT CS
Response_reg 1 vlivdDeplvrellrqalekegyeevaeaddgeealellkekd..pDlillDiempgmdGlellkeireeepklpiivvtahgeeedalealkaG 93
vl+vdD+p+ +++l+++l+ y ev+ + +e al ll+e++ +D+++ D+ mp+m+G++ll++i e +lp+i+++a + ++ + + + G
XP_010101901.1 26 VLVVDDDPTCLMILEKMLRTCLY-EVTKCNRAEHALSLLRENKngFDIVISDVHMPDMNGFKLLEHIGLEM-DLPVIMMSADDGKDVVMKGVTHG 118
89*********************.***************999999**********************6654.8********************** PP
ESEEEESS--HHHHHH CS
Response_reg 94 akdflsKpfdpeelvk 109
a d+l Kp+ +e+l +
XP_010101901.1 119 ACDYLIKPVRIEALKN 134
************9986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PIRSF | PIRSF036392 | 2.5E-180 | 6 | 646 | IPR017053 | Response regulator B-type, plant |
| Gene3D | G3DSA:3.40.50.2300 | 6.2E-42 | 23 | 167 | No hit | No description |
| SuperFamily | SSF52172 | 2.99E-35 | 23 | 147 | IPR011006 | CheY-like superfamily |
| SMART | SM00448 | 3.2E-30 | 24 | 136 | IPR001789 | Signal transduction response regulator, receiver domain |
| PROSITE profile | PS50110 | 42.319 | 25 | 140 | IPR001789 | Signal transduction response regulator, receiver domain |
| Pfam | PF00072 | 5.0E-23 | 26 | 134 | IPR001789 | Signal transduction response regulator, receiver domain |
| CDD | cd00156 | 1.94E-27 | 27 | 139 | No hit | No description |
| SuperFamily | SSF46689 | 3.4E-20 | 201 | 261 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 11.048 | 201 | 260 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.3E-30 | 203 | 262 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 9.3E-25 | 204 | 257 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 9.3E-8 | 206 | 256 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009873 | Biological Process | ethylene-activated signaling pathway | ||||
| GO:0010082 | Biological Process | regulation of root meristem growth | ||||
| GO:0010119 | Biological Process | regulation of stomatal movement | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0071368 | Biological Process | cellular response to cytokinin stimulus | ||||
| GO:0080113 | Biological Process | regulation of seed growth | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 654 aa Download sequence Send to blast |
MATSTSGAAW KSGDVVSDQF PAGLRVLVVD DDPTCLMILE KMLRTCLYEV TKCNRAEHAL 60 SLLRENKNGF DIVISDVHMP DMNGFKLLEH IGLEMDLPVI MMSADDGKDV VMKGVTHGAC 120 DYLIKPVRIE ALKNIWQHVV RKKKNEWREL EQSGSVEEGD RSAKPSEDAD YSSSANEGNW 180 KSSKKRRDED EEAEERDDTS TLKKPRVVWS VELHQQFVAA VNQLGIDKAV PKKILELMNV 240 PGLTRENVAS HLQKYRLYLR RLSGQHQTSM NNSFINPQEA SFGAMTSLNG IDLQTLAVAG 300 QIPAQSLATL QAAGLGRSTA KPGIPLPLVD QRNLFSFENP KLRFGEGQQQ QQQHLGTSKP 360 VNLLHGIPTS MEPKQLTSLH QSAQSLGMNM QVNTHGAQNS SLLMQMNQPQ SRGQILNDTA 420 SSHVPRLPTT QTIVSTGLAS GVLGRNGITN NGRGAGYNPV MQNSSMLNFP NGFPLGSAST 480 MSGLPSQGSF QEEVTSEVKG SVSFAASYDI FNDLNQHKSN DWDIQNVGLT FDASQQSNSI 540 PSGSLDLVHQ GFSGSQRNMP VRNQSNVGKP MYSIGDIGHG NAQNTSQHLN TLLGDNSIRI 600 KTERNTDVLA DNGFFPEQFG QDDLMSALLK QQEGITPAEN QFDFDGYSMD NIPV |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 3e-22 | 203 | 263 | 4 | 64 | ARR10-B |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specifically to the DNA sequence 5'-[AG]GATT-3'. Functions as a response regulator involved in His-to-Asp phosphorelay signal transduction system. Phosphorylation of the Asp residue in the receiver domain activates the ability of the protein to promote the transcription of target genes. Could directly activate some type-A response regulators in response to cytokinins. Involved in the expression of nuclear genes for components of mitochondrial complex I. Promotes cytokinin-mediated leaf longevity. Involved in the ethylene signaling pathway in an ETR1-dependent manner and in the cytokinin signaling pathway. {ECO:0000269|PubMed:11370868, ECO:0000269|PubMed:11574878, ECO:0000269|PubMed:15282545, ECO:0000269|PubMed:16407152}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00054 | PBM | Transfer from AT4G16110 | Download |
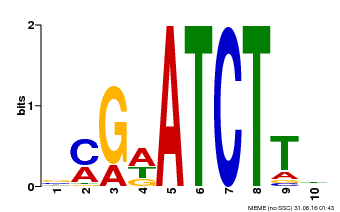
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_010101901.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010101901.1 | 0.0 | two-component response regulator ARR2 isoform X2 | ||||
| Swissprot | Q9ZWJ9 | 0.0 | ARR2_ARATH; Two-component response regulator ARR2 | ||||
| TrEMBL | W9RS74 | 0.0 | W9RS74_9ROSA; Two-component response regulator | ||||
| STRING | XP_010101901.1 | 0.0 | (Morus notabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF3958 | 34 | 58 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G16110.1 | 0.0 | response regulator 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Entrez Gene | 21388359 |



