 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010091413.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Moraceae; Morus
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 1026aa MW: 113088 Da PI: 6.4613 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 129.1 | 1.7e-40 | 172 | 249 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cqve+C adls+ak+yhrrhkvCe+hska ++lv ++ qrfCqqCsrfh l+efDe+krsCrrrLa+hn+rrrk+++
XP_010091413.1 172 VCQVEDCGADLSSAKDYHRRHKVCEMHSKACKALVGNVLQRFCQQCSRFHVLQEFDEGKRSCRRRLAGHNKRRRKTNP 249
6**************************************************************************976 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 1.7E-32 | 165 | 234 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 31.958 | 170 | 247 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 1.44E-37 | 171 | 252 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 6.2E-29 | 173 | 246 | IPR004333 | Transcription factor, SBP-box |
| CDD | cd00204 | 1.07E-5 | 770 | 913 | No hit | No description |
| SuperFamily | SSF48403 | 1.61E-5 | 797 | 913 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 5.0E-5 | 799 | 913 | IPR020683 | Ankyrin repeat-containing domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0016021 | Cellular Component | integral component of membrane | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1026 aa Download sequence Send to blast |
MEARFGGEAH HFYGMSTADL PKRANLEWDL NHWKWDGDLF IASSVVNPVV GVGVGPSSHA 60 MASSSSRQFF PLGSGAGGSS NSSSSCSEGG NLGMIEKGKR ELMVEKRRRV NVVEEEDNLN 120 DGDEAGTLTL KLGGGGRVYN QTSEREVGVN NWEGTSGKKT KLAAGGSSSR AVCQVEDCGA 180 DLSSAKDYHR RHKVCEMHSK ACKALVGNVL QRFCQQCSRF HVLQEFDEGK RSCRRRLAGH 240 NKRRRKTNPD PVVNGSSLND DQTSGYLLIS LLRILSNMHS NRSDQSHQTT DQDLLSHLLR 300 SLASQTSDHG GKNIAGLLQE PQKLLNEGTS VGNSDVVSTF IANSSQGPPR PIKQHQTVSV 360 SEIPQQGVHL HNANGGSIQA TSSIKPSILN SPPSYSEARD GTAGQIKMNN FDLNDIYIDS 420 DDSVEDPERS PPTTNAVTSS LDCPSWVQQD SHQSSPPQTS GNSDSASAQS PSSSSGEAQS 480 RTDRIVFKLF GKEPNDFPLV LRAQILDWLS HSPSEIESYI RPGCIILTIY LRQSETAWEE 540 LCDDLSSSLS RLLDVSDDSF WRSGWIFIRA QHQIAFIYNG QVVVDTSLPL RSSNYSKIVS 600 VEPIAVPASE RAQFSVRGIN LVRPTTRLFC ALEGKYLVQE ATHELMESVD NVEHDEQCIN 660 FSCPIPVTNG RGFIEIEDQG LGSSFFPFIV AEEDVCSEIR VLESSLEHGR TGKPDTYNQA 720 VDFIHEMGWL LHRSQLRSRL GHLDPNADPF PLKRFKWIME FSMDHDWSAV VRKLLDILHD 780 GNVGAGDDHS ISLALSEMGL LHRAVRRNSR PLVEVLLKYV PKNLSNNSES EDKAVSNEVN 840 KGFLFRPDVI GPASLTPLHI AAGKDGSEDV LDALTNDPGM VGIEAWKSAH DSTGSTPEDY 900 ARLRGHYSYI RLIQRKINKR PASGHVVVDI PSNLNDCSTS QKQNEPVSSF QIGRTELRRN 960 QHPCRLCDRK LVYGTTSSSV VYRPAMLSMV AIAAVCVCVA LLFKSSPEVL YVFQPFRWER 1020 LEYGTS |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 1e-30 | 166 | 246 | 4 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 105 | 109 | KRRRV |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' of AP1 promoter. Binds specifically to the 5'-GTAC-3' core sequence. {ECO:0000269|PubMed:10524240, ECO:0000269|PubMed:16095614}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00633 | PBM | Transfer from PK06791.1 | Download |
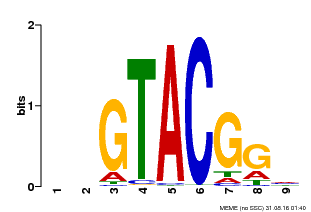
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_010091413.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010091413.1 | 0.0 | squamosa promoter-binding-like protein 1 | ||||
| Refseq | XP_024018150.1 | 0.0 | squamosa promoter-binding-like protein 1 | ||||
| Swissprot | Q9SMX9 | 0.0 | SPL1_ARATH; Squamosa promoter-binding-like protein 1 | ||||
| TrEMBL | W9QN82 | 0.0 | W9QN82_9ROSA; Squamosa promoter-binding-like protein 1 | ||||
| STRING | XP_010091413.1 | 0.0 | (Morus notabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1539 | 34 | 95 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Entrez Gene | 21397031 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



