 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Migut.O00397.1.p | ||||||||
| Common Name | LOC105970489, MIMGU_mgv1a021370mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Phrymaceae; Erythranthe
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 930aa MW: 103143 Da PI: 7.3054 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 129.1 | 1.6e-40 | 122 | 199 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cqve+C+adls+ak+yhrrhkvC+ hsk++++lv +++qrfCqqCsrfh l+efDe+krsCrrrLa+hn+rrrk+++
Migut.O00397.1.p 122 VCQVEDCRADLSNAKDYHRRHKVCDLHSKSTSALVGDVMQRFCQQCSRFHVLQEFDEGKRSCRRRLAGHNKRRRKTHP 199
6**************************************************************************976 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 1.8E-32 | 118 | 184 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 31.727 | 120 | 197 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 7.32E-37 | 122 | 200 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 7.5E-29 | 123 | 196 | IPR004333 | Transcription factor, SBP-box |
| CDD | cd00204 | 1.88E-5 | 684 | 818 | No hit | No description |
| SuperFamily | SSF48403 | 5.51E-5 | 716 | 819 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 1.9E-5 | 717 | 817 | IPR020683 | Ankyrin repeat-containing domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0016021 | Cellular Component | integral component of membrane | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 930 aa Download sequence Send to blast |
MESKIGGEKI HKSMEWDSNE WRWDGALLVA TPLNSVPSDC RSRQLGSDEL VLLGDGNGNG 60 REKRDFEKRA RGVFEMEENE NEDVGLPNLK LGGQVYPIAE MEGKSGKKTK VSNALPMSSP 120 SVCQVEDCRA DLSNAKDYHR RHKVCDLHSK STSALVGDVM QRFCQQCSRF HVLQEFDEGK 180 RSCRRRLAGH NKRRRKTHPE NDEPGSNYLL ISLLRILSNI HSNSSDQIQD QDLVSHLLKN 240 LAHLTGTNPA GVLPVSQNVG TSLGTALKGL SAPSGPGMTI PASDLTEKRT LIGGVSHAST 300 SESPLPFRTT SSDLFKEKDS NTGVGRTKLS NIDLNCAYDG SQDCMEDMPN TSHLNKTSPG 360 GSSWLCKDSQ RCGPPQNSGN SASTSSQSPS TSSGEAQSRT DRIVFKLFGK DPSDFPLLLR 420 KQILDWLSNS PTDIESYIRP GCIILTIYLH MEKSSWDELY CNLTSSLLRL LNSSTDSFWR 480 TGWIYTRVHH HVTFMYNGQV VLDTPLPVRN HQSCRISSIK PIAVTVSEGV HFVVKGFNLS 540 RSTSRLLCAL EGKYLVQENC GDMIGRADSF IEHNQIQSLT FSCAVPNIVG RGFIEIEDHG 600 LSSSFFPFIV AEKDVCSEIC TLESVIEDAN EIQVRNEALD FIHEMGWLLQ RNRLKSRLGD 660 GDLFPFERFR RLTEFSVDHD WCAVVKKLLR ILFDDGTVDL GPQNSNIVAL LNDVGLVHRA 720 VRRKCSSMVQ FLLNETNPLA DDGGGAHLYL FRPDAAGPGG LTPLHIAASL DGCENVLDAL 780 TEDPGSVGIE EWKRGRDSSG LTAHDYACIR GQYSYINIVQ RKVDKKSAVV GVHIGDSSSS 840 RGEVVLSVSV EKTMEIERRR RRCGECEERI MRYGNRSTRG RVRIYRPAML SLVGIAAVCV 900 CTALLFKSSP EVLFSFRPFR WDQLKYGSQ* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 8e-30 | 122 | 196 | 10 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 191 | 197 | KRRRKTH |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000269|PubMed:16554053}. | |||||
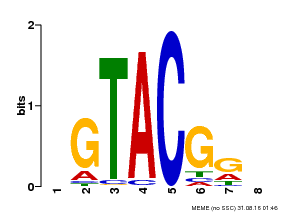
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00060 | PBM | Transfer from AT3G60030 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Migut.O00397.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KT717966 | 1e-179 | KT717966.1 Erythranthe guttata squamosa promoter-binding protein 3 mRNA, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012850775.1 | 0.0 | PREDICTED: squamosa promoter-binding-like protein 12 | ||||
| Refseq | XP_012850776.1 | 0.0 | PREDICTED: squamosa promoter-binding-like protein 12 | ||||
| Swissprot | Q9S7P5 | 0.0 | SPL12_ARATH; Squamosa promoter-binding-like protein 12 | ||||
| TrEMBL | A0A022QE34 | 0.0 | A0A022QE34_ERYGU; Uncharacterized protein | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA1871 | 24 | 66 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G60030.1 | 0.0 | squamosa promoter-binding protein-like 12 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Migut.O00397.1.p |
| Entrez Gene | 105970489 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



