 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Migut.M00094.2.p | ||||||||
| Common Name | LOC105951952, MIMGU_mgv1a001241mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Phrymaceae; Erythranthe
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 847aa MW: 96723.2 Da PI: 6.3303 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 87.8 | 1.6e-27 | 86 | 188 | 2 | 91 |
FAR1 2 fYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkk.............tekerrtraetrtgCkaklkvkkekdgkwevtkl 81
fY+eYA++vGFs+ +++s++sk+++e+++++f Cs++g+++e +k+ +e++ +ra +t+Cka+++vk+++dgkw ++++
Migut.M00094.2.p 86 FYQEYARSVGFSTAIQNSRRSKTSREFIDAKFACSRYGTKREYEKSlnrprsrqganqdAENATGRRACAKTDCKASMHVKRRSDGKWIIHRF 178
9*****************************************999999***999998776777779*************************** PP
FAR1 82 eleHnHelap 91
e+eHnHel p
Migut.M00094.2.p 179 EKEHNHELLP 188
*******975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 1.1E-25 | 86 | 188 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 3.6E-28 | 286 | 378 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 9.507 | 566 | 602 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 1.7E-4 | 574 | 599 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 1.1E-7 | 577 | 604 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0010218 | Biological Process | response to far red light | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 847 aa Download sequence Send to blast |
MDIDLRLHSG EQDKEVEETN GIVVMLDGEE KPLNIEGSVE DIEEKLHIED EEEVGSPLND 60 IDFKDVTILE PLPGMEFGSH GDAYAFYQEY ARSVGFSTAI QNSRRSKTSR EFIDAKFACS 120 RYGTKREYEK SLNRPRSRQG ANQDAENATG RRACAKTDCK ASMHVKRRSD GKWIIHRFEK 180 EHNHELLPAQ AVSEQTRRMY AAMARQFAEY KSAVGLKHDP RSQFEKARNT ALDAGDVNIL 240 LEFFVQMQRL NSNFFYAVDA GEDQRLKNFL WVDAKSRHDY ASFSDVVSFD TSYVRNKYKM 300 PLALFVGVNQ HYQFMLLGCA LICDENAGTY SWVMQTWLKA MGGQAPKIII TDQDEAMKSV 360 ISDVFPSALH FFCLWNITGK VSETLSHVIK QNEKFMLKFE KCVYRSWTDD EFEKRWHKLV 420 ERFELQENEL IQSLYEDREK WVPNFMKDGF LAGMSTGVRS ESVNSFFDKY VHKKTTVQEF 480 LKQYETILQD RYEEEAKASS DTWNKPPVLK SPSPFEKHVA GLYTHAVFRK FQVEVLGAVA 540 CIPKREEQVD ATVTFKVQDF ETSREFVVTL NEMKSEISCI CRLFEFKGFL CRHAMIVLQI 600 CGISTIPMQY ILKRWTKDAK SRYSMGEGSE MAQTRLQRYN DLCQKAIKLG EEGSLSQESY 660 NMTLRALEDA FENCLNANNC NRNLLEAGPS ASPGILCIEE DIQSGSLSKT NKKKNNTTTK 720 KRKVNMEQDV ITVGATESMQ QMEKLSSSRP VNLDGFFGPQ QSVQGMVQLN LMGPARDNYY 780 GNQQTIQGLG QLNSIAPTHD GYYGTQPAIH GLGQMDFFRT PSFGYGIRED PNVRPAQLHD 840 DATRHA* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. When associated with PHYA, protects it from being recognized and degraded by the COP1/SPA complex. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:17012604, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:18715961, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00078 | ChIP-seq | Transfer from AT3G22170 | Download |
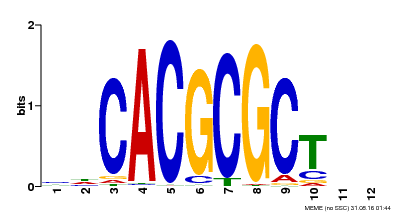
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Migut.M00094.2.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. Up-regulated by white light. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012830894.1 | 0.0 | PREDICTED: protein FAR-RED ELONGATED HYPOCOTYL 3 isoform X2 | ||||
| Swissprot | Q9LIE5 | 0.0 | FHY3_ARATH; Protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| TrEMBL | A0A022RST6 | 0.0 | A0A022RST6_ERYGU; Uncharacterized protein | ||||
| STRING | Migut.M00094.1.p | 0.0 | (Erythranthe guttata) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G22170.2 | 0.0 | far-red elongated hypocotyls 3 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Migut.M00094.2.p |
| Entrez Gene | 105951952 |



