 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Migut.L01319.1.p | ||||||||
| Common Name | LOC105967482, MIMGU_mgv1a021435mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Phrymaceae; Erythranthe
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 266aa MW: 30502 Da PI: 6.2605 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 59.2 | 6.7e-19 | 67 | 120 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k+++++ eq+++Le+ Fe+++++ e++ +LA+ lgL+ rq+ +WFqNrRa++k
Migut.L01319.1.p 67 KKRRLNIEQVRTLEKSFESCNKLEPERKIQLARALGLQPRQIAIWFQNRRARWK 120
4557899**********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 127.1 | 7.5e-41 | 66 | 158 | 1 | 93 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLreelke 93
ekkrrl+ eqv++LE+sFe+ +kLeperK +lar+Lglqprq+a+WFqnrRAR+ktkqlEkdy+ Lkr++da+k en++L + +++L++e+k+
Migut.L01319.1.p 66 EKKRRLNIEQVRTLEKSFESCNKLEPERKIQLARALGLQPRQIAIWFQNRRARWKTKQLEKDYDLLKRQFDAVKLENDALLTLNQKLHAEIKA 158
69***************************************************************************************9986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.13E-19 | 58 | 124 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 16.925 | 62 | 122 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.3E-19 | 65 | 126 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 5.25E-17 | 67 | 123 | No hit | No description |
| Pfam | PF00046 | 3.0E-16 | 67 | 120 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 1.4E-19 | 69 | 129 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 1.5E-5 | 93 | 102 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 97 | 120 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 1.5E-5 | 102 | 118 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 5.5E-13 | 122 | 162 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009744 | Biological Process | response to sucrose | ||||
| GO:0048826 | Biological Process | cotyledon morphogenesis | ||||
| GO:0080022 | Biological Process | primary root development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 266 aa Download sequence Send to blast |
MGFFHSNFML QINPQQEDDH HTTQFHGVAS SLLGKRPLDE DYTNNNNIIN HCGDDELSDD 60 GSNIGEKKRR LNIEQVRTLE KSFESCNKLE PERKIQLARA LGLQPRQIAI WFQNRRARWK 120 TKQLEKDYDL LKRQFDAVKL ENDALLTLNQ KLHAEIKALK KREPTESINL NKETEGSSSN 180 NRSENSSEMK NIALSDINYY SSRNDASIDG PLTAAIIRPL FPSRVTDNIQ TAVKEESRSS 240 SLCNLFCSMD DQTGFWPWLD QPNFN* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 114 | 122 | RRARWKTKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that may act in the sucrose-signaling pathway. {ECO:0000269|PubMed:11292072}. | |||||
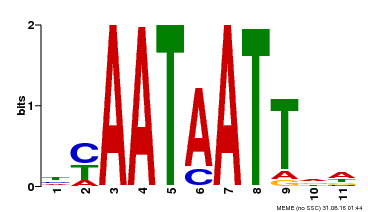
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00225 | DAP | Transfer from AT1G69780 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Migut.L01319.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012847536.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ATHB-13 | ||||
| Swissprot | Q8LC03 | 9e-90 | ATB13_ARATH; Homeobox-leucine zipper protein ATHB-13 | ||||
| TrEMBL | A0A022QL88 | 0.0 | A0A022QL88_ERYGU; Uncharacterized protein | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA848 | 24 | 98 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G69780.1 | 6e-85 | HD-ZIP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Migut.L01319.1.p |
| Entrez Gene | 105967482 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



