 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Migut.K01320.1.p | ||||||||
| Common Name | LOC105960954, MIMGU_mgv1a008126mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Phrymaceae; Erythranthe
|
||||||||
| Family | Dof | ||||||||
| Protein Properties | Length: 385aa MW: 41882.6 Da PI: 4.986 | ||||||||
| Description | Dof family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-Dof | 123.5 | 7.2e-39 | 101 | 160 | 2 | 61 |
zf-Dof 2 kekalkcprCdstntkfCyynnyslsqPryfCkaCrryWtkGGalrnvPvGggrrknkks 61
++k+l+cprC+s++tkfCy+nny+++qPr+fCk+C+ryWt+GG++rnvPvG+grr+nk++
Migut.K01320.1.p 101 PDKILPCPRCNSMETKFCYFNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRRNKNV 160
7899*****************************************************975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| ProDom | PD007478 | 1.0E-30 | 88 | 158 | IPR003851 | Zinc finger, Dof-type |
| Pfam | PF02701 | 1.8E-31 | 103 | 159 | IPR003851 | Zinc finger, Dof-type |
| PROSITE profile | PS50884 | 28.779 | 105 | 159 | IPR003851 | Zinc finger, Dof-type |
| PROSITE pattern | PS01361 | 0 | 107 | 143 | IPR003851 | Zinc finger, Dof-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 385 aa Download sequence Send to blast |
MAEVKDPKIK LFGKTIELPE TSAAAAAAEE NVVAAQSCEA AGDDSSNQDP PCSSNSNGDA 60 EDQESLKIHS GPKQDDVKEK DQTEQDQSET ENSDEKALKK PDKILPCPRC NSMETKFCYF 120 NNYNVNQPRH FCKNCQRYWT AGGTMRNVPV GAGRRRNKNV GPTQLRHQNV PPEALQTLRP 180 NPNPSNPTLC ESMVSVLNIA DKKIPNKGDE PSKGETGLHE KLPNCPPYFP GAAWPNYPWQ 240 VVPLYPTPPP PFWGCTVQPL QPWSVPWVIP QMPSSSGGPN SPTLGKRNLD EEEIQESEGK 300 SEKLLWVPKT LRIDDPGEAA KSSIWATLGI KNDGGVDSSG GGLFEPFERG TKSDEKVPEA 360 STTALQANPA AMSRSVSFRE SSSS* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
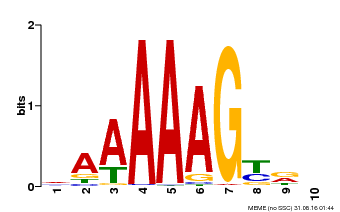
| UniProt | Transcription factor that binds specifically to a 5'-AA[AG]G-3' consensus core sequence (By similarity). Regulates a photoperiodic flowering response. Transcriptional repressor of 'CONSTANS' expression. The stability of CDF2 is controlled by 'GIGANTEA' and redundantly by ADO3, ADO2 and/or ADO1. {ECO:0000250, ECO:0000269|PubMed:19619493}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00046 | PBM | Transfer from AT5G39660 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Migut.K01320.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Circadian-regulation. Highly expressed at the beginning of the light period, then decreases, reaching a minimum between 16 and 29 hours after dawn before rising again at the end of the day. Regulated at the protein level by ADO3 and GI pos-transcriptionally. {ECO:0000269|PubMed:19619493}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012840628.1 | 0.0 | PREDICTED: cyclic dof factor 2-like | ||||
| Swissprot | Q93ZL5 | 3e-86 | CDF2_ARATH; Cyclic dof factor 2 | ||||
| TrEMBL | A0A022R705 | 0.0 | A0A022R705_ERYGU; Uncharacterized protein | ||||
| STRING | Migut.K01320.1.p | 0.0 | (Erythranthe guttata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA532 | 24 | 122 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G39660.2 | 3e-69 | cycling DOF factor 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Migut.K01320.1.p |
| Entrez Gene | 105960954 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



