 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Migut.G00007.1.p | ||||||||
| Common Name | MIMGU_mgv1a000988mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Phrymaceae; Erythranthe
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 939aa MW: 104238 Da PI: 7.5763 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 129.1 | 1.6e-40 | 129 | 206 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cqve+C+adls+ak+yhrrhkvC+ hsk++++lv +++qrfCqqCsrfh l+efDe+krsCrrrLa+hn+rrrk+++
Migut.G00007.1.p 129 VCQVEDCRADLSNAKDYHRRHKVCDLHSKSTSALVGDVMQRFCQQCSRFHVLQEFDEGKRSCRRRLAGHNKRRRKTHP 206
6**************************************************************************976 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 6.3E-32 | 124 | 191 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 31.684 | 127 | 204 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 1.83E-36 | 129 | 207 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 7.6E-29 | 130 | 203 | IPR004333 | Transcription factor, SBP-box |
| CDD | cd00204 | 2.98E-4 | 691 | 825 | No hit | No description |
| Gene3D | G3DSA:1.25.40.20 | 3.9E-4 | 724 | 824 | IPR020683 | Ankyrin repeat-containing domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0016021 | Cellular Component | integral component of membrane | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 939 aa Download sequence Send to blast |
MESKIGGEKI HKSMEWDSNE WRWDGALFVA TPLNSVPSDC RSRQLGSDEL VLLGDGNGNG 60 REKRELEKRT RGVLEMEENE SEDVGLPNLK LGGQVYPIAE RDMYEMEGKS GKKTKVSSAL 120 PMSTSSRSVC QVEDCRADLS NAKDYHRRHK VCDLHSKSTS ALVGDVMQRF CQQCSRFHVL 180 QEFDEGKRSC RRRLAGHNKR RRKTHPENDE PGTNYLLISL LRILSNIHSN SSDQIQDQDL 240 VSHLLKNLAH LTGTNPAGVL PISQNVGTSL GTALKGLSAP SGPGVTIPAS DLTEKRTLIG 300 GVSHASTSES PLPFRTTSSD LFKEKDSNTG VGRTKLSNID LNCAYDGSQD CMEDMPNTSH 360 LNKTSPGGSS WLCKDSQRCG PPQNSGNSAS TSSQSPSTSS GEAQSRTDRI VFKLFGKDPS 420 DFPLLLRKQI LDWLSNSPTD IESYIRPGCI ILTIYLRMEK SSWDELYCNL TSSLLRLLNS 480 STDSFWRTGW IYTRVHHHVT FMYNGQVVLD TPLPVRNHQS CRISSIKPIA VTVSEGVHFV 540 VKGFNLSRST SRLLCALEGK YLVQENCADM TGRADSLTEH DQIQSLTFSC AVPNIVGRGF 600 IEIEDHGLSS SFFPFIVAEK DVCSEICSLE SVIEDANEIQ VRNEALDFIH EMGWLLQRNR 660 LKSRLGDGDL FPFERFRRLT EFSIDHDWCA VVKKLLRILF DDGTVDLGPH NSNIVALLND 720 VGLVHRAVRR KCSSMVRFLL NEKNPLADGG GGAHLYLFRP DAAGPGGLTP LHIAASLDGC 780 ENVLDALTED PGSVGIEEWK RGRDSSVLTA HDYACIRGQY SYINIVQRKV DKKSTVVGVG 840 VDIGDSSSSR GEVVLSVSVD KKMEIERRRR RCGECEERIM RYGNRSTRGR VRIYRPAMLS 900 LVGIAAVCVC TALLFKSSPE VLFSFRPFRW DQLKYGSQ* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 7e-30 | 129 | 203 | 10 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 198 | 204 | KRRRKTH |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000269|PubMed:16554053}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00633 | PBM | Transfer from PK06791.1 | Download |
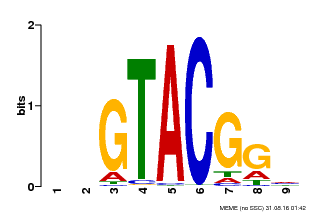
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Migut.G00007.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KT717966 | 1e-162 | KT717966.1 Erythranthe guttata squamosa promoter-binding protein 3 mRNA, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012842609.1 | 0.0 | PREDICTED: squamosa promoter-binding-like protein 12 | ||||
| Swissprot | Q9S7P5 | 0.0 | SPL12_ARATH; Squamosa promoter-binding-like protein 12 | ||||
| TrEMBL | A0A022R2N5 | 0.0 | A0A022R2N5_ERYGU; Uncharacterized protein | ||||
| STRING | Migut.G00007.1.p | 0.0 | (Erythranthe guttata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA1871 | 24 | 66 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G60030.1 | 0.0 | squamosa promoter-binding protein-like 12 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Migut.G00007.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



