 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Migut.D00401.1.p | ||||||||
| Common Name | LOC105952451, MIMGU_mgv1a005352mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Phrymaceae; Erythranthe
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 488aa MW: 53949.4 Da PI: 6.334 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 40 | 7.2e-13 | 338 | 384 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCC.C...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPk.askapskKlsKaeiLekAveYIksLq 55
+h e+Er+RR+++N++f Lr ++P+ + K++Ka+ L A+ YI +Lq
Migut.D00401.1.p 338 NHVEAERQRREKLNQRFYALRAVVPNiS------KMDKASLLGDAIAYITDLQ 384
799***********************66......*****************99 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 6.3E-43 | 49 | 229 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| PROSITE profile | PS50888 | 16.66 | 334 | 383 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 3.27E-18 | 334 | 399 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 1.28E-14 | 337 | 388 | No hit | No description |
| Gene3D | G3DSA:4.10.280.10 | 7.2E-18 | 338 | 402 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 2.2E-10 | 338 | 384 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 7.1E-16 | 340 | 389 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 488 aa Download sequence Send to blast |
MGNKFWLNGE DKAVVESVLG SEATEYFAWS ASNNVLSEFS ASGGDLGVQQ TLCKIVDGSD 60 WTYAIYWHVS RSKSGKSALI WGDGHCREPK EGEYDTGNFS EDGKKPVEGD CRKRVLQKIH 120 ACFGGSEDDN VAAKLDQVSH VEMFYLTSMY FAFPFDKPSI PSQSFNSGRT IWVSDMKSCL 180 ERYQSRSYLA KLAGFETLVY IPSKSGVIEI GSRKSIPENQ SIIQSAKSII VKVNSTHAKP 240 FLPKVFGQEL NLGGSKSSTI SISFSPKVED DSVLSSESYD LQVYGNSSNG HRTDDIDAKI 300 YPHLNMDQIA ENSFVLSDDQ RPRKRGRKPA NGREEALNHV EAERQRREKL NQRFYALRAV 360 VPNISKMDKA SLLGDAIAYI TDLQTKVRVM EAEKGVVNNN NNKQHESNIP DIDFLERHED 420 SVLRVSCPLE SHPVSGVVRA LRENQVAAHE SVISINEGGE VVHTFTIQTP GGGAEELKDK 480 LAAALLK* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5gnj_A | 1e-28 | 332 | 399 | 2 | 69 | Transcription factor MYC2 |
| 5gnj_B | 1e-28 | 332 | 399 | 2 | 69 | Transcription factor MYC2 |
| 5gnj_E | 1e-28 | 332 | 399 | 2 | 69 | Transcription factor MYC2 |
| 5gnj_F | 1e-28 | 332 | 399 | 2 | 69 | Transcription factor MYC2 |
| 5gnj_G | 1e-28 | 332 | 399 | 2 | 69 | Transcription factor MYC2 |
| 5gnj_I | 1e-28 | 332 | 399 | 2 | 69 | Transcription factor MYC2 |
| 5gnj_M | 1e-28 | 332 | 399 | 2 | 69 | Transcription factor MYC2 |
| 5gnj_N | 1e-28 | 332 | 399 | 2 | 69 | Transcription factor MYC2 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that negatively regulates jasmonate (JA) signaling (PubMed:30610166). Negatively regulates JA-dependent response to wounding, JA-induced expression of defense genes, JA-dependent responses against herbivorous insects, and JA-dependent resistance against Botrytis cinerea infection (PubMed:30610166). Plays a positive role in resistance against the bacterial pathogen Pseudomonas syringae pv tomato DC3000 (PubMed:30610166). {ECO:0000269|PubMed:30610166}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00099 | PBM | Transfer from AT4G16430 | Download |
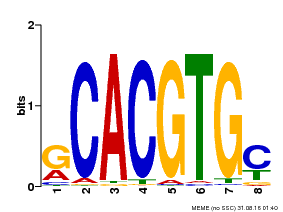
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Migut.D00401.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by wounding, feeding with herbivorous insects, infection with the fungal pathogen Botrytis cinerea and infection with the bacterial pathogen Pseudomonas syringae pv tomato DC3000. {ECO:0000269|PubMed:30610166}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012831462.1 | 0.0 | PREDICTED: transcription factor bHLH3 | ||||
| Swissprot | A0A3Q7H216 | 0.0 | MTB3_SOLLC; Transcription factor MTB3 | ||||
| TrEMBL | A0A022S146 | 0.0 | A0A022S146_ERYGU; Uncharacterized protein | ||||
| STRING | Migut.D00401.1.p | 0.0 | (Erythranthe guttata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA6669 | 24 | 33 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G16430.1 | 1e-151 | bHLH family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Migut.D00401.1.p |
| Entrez Gene | 105952451 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



