 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Migut.C00805.1.p | ||||||||
| Common Name | MIMGU_mgv1a026529mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Phrymaceae; Erythranthe
|
||||||||
| Family | E2F/DP | ||||||||
| Protein Properties | Length: 414aa MW: 45611.5 Da PI: 7.9202 | ||||||||
| Description | E2F/DP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | E2F_TDP | 86.1 | 2.6e-27 | 151 | 216 | 1 | 71 |
E2F_TDP 1 rkeksLrlltqkflkllekseegivtlnevakeLvsedvknkrRRiYDilNVLealnliekkekneirwkg 71
r+++sL+llt+kf+kl++++++g+++ln a+ L +v ++RRiYDi+NVLe+++liek+ kn+irwkg
Migut.C00805.1.p 151 RYDSSLGLLTKKFIKLIQEAKDGTLDLNRTADVL---QV--QKRRIYDITNVLEGIGLIEKTTKNHIRWKG 216
6899******************************...99..****************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.10 | 3.5E-28 | 150 | 217 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| SMART | SM01372 | 4.9E-35 | 151 | 216 | IPR003316 | E2F/DP family, winged-helix DNA-binding domain |
| SuperFamily | SSF46785 | 1.5E-17 | 151 | 214 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| Pfam | PF02319 | 1.6E-25 | 153 | 216 | IPR003316 | E2F/DP family, winged-helix DNA-binding domain |
| CDD | cd14660 | 9.98E-36 | 227 | 333 | No hit | No description |
| SuperFamily | SSF144074 | 1.57E-25 | 230 | 334 | No hit | No description |
| Pfam | PF16421 | 1.1E-23 | 232 | 331 | IPR032198 | E2F transcription factor, CC-MB domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000902 | Biological Process | cell morphogenesis | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0042023 | Biological Process | DNA endoreduplication | ||||
| GO:0051782 | Biological Process | negative regulation of cell division | ||||
| GO:0005667 | Cellular Component | transcription factor complex | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| GO:0046982 | Molecular Function | protein heterodimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 414 aa Download sequence Send to blast |
MSTSGENLDL NLSLNLPLSR MQLENQAAPS LGRVRSNRFA CLPPPSNTAS SCGLASDKFA 60 ANYNNGVTAQ QSRTSVQTPP KNDPSIEGKI CGNTSIQVPK QEQTAYNGFP GPAYMMARSS 120 KKAKSVKHDK NAVKEAKLEP GDNLNLTGCC RYDSSLGLLT KKFIKLIQEA KDGTLDLNRT 180 ADVLQVQKRR IYDITNVLEG IGLIEKTTKN HIRWKGFGTV GSQKLDDGVS YLKAEVEHLS 240 AEECRLDNQI RDKQESLREL ISNQNCKKNL FLTKDDIMSL PSLRNQTVIA VEAPRASSIA 300 VPDPDEDIGF CQKQYSLMIR STTGPIAVYL LSKNEKKNKE ISVKRAKLMD PFTWTNSSRV 360 DDAELSLSPD TSTSSKGCGF QKIVPIDSGI DDDYWLRSEH EVSATDLWGT EEL* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1cf7_A | 2e-25 | 150 | 218 | 6 | 75 | PROTEIN (TRANSCRIPTION FACTOR E2F-4) |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Involved in transcriptional repression. May act by repressing E2F-regulated genes in mature differentiated cells, but is not an antagonist of E2FA. Restricts cell division and is involved in the coordination between cell proliferation and endoreduplication during development. May play a role during the transition from skotomorphogenesis to photomorphogenesis. Regulated by phosphorylation-dependent proteolysis via the protein-ubiquitin ligase SCF(SKP2A) complex. {ECO:0000269|PubMed:11786543, ECO:0000269|PubMed:11891240, ECO:0000269|PubMed:12468727, ECO:0000269|PubMed:16920782, ECO:0000269|PubMed:19662336}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00190 | DAP | Transfer from AT1G47870 | Download |
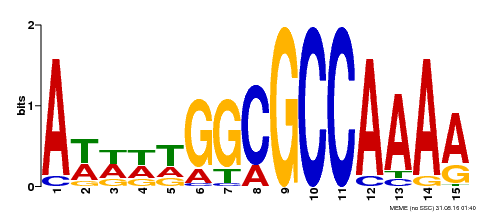
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Migut.C00805.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated by light. {ECO:0000269|PubMed:12468727, ECO:0000269|PubMed:18424613}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012835754.1 | 0.0 | PREDICTED: transcription factor E2FC | ||||
| Swissprot | Q9FV70 | 4e-84 | E2FC_ARATH; Transcription factor E2FC | ||||
| TrEMBL | A0A022RGC2 | 0.0 | A0A022RGC2_ERYGU; Uncharacterized protein | ||||
| STRING | Migut.C00805.1.p | 0.0 | (Erythranthe guttata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA12625 | 20 | 22 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G47870.2 | 2e-86 | E2F/DP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Migut.C00805.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



