 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Migut.C00339.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Phrymaceae; Erythranthe
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 1050aa MW: 115496 Da PI: 8.1228 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 128.1 | 3.3e-40 | 167 | 244 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cqv++C +dls ak+yhrrhkvCe+hska ++lv +++qrfCqqCsrfh lsefDe+krsCrrrLa+hn+rrrk+q+
Migut.C00339.1.p 167 VCQVDNCVKDLSAAKDYHRRHKVCEFHSKAGNALVGKQMQRFCQQCSRFHPLSEFDEGKRSCRRRLAGHNRRRRKTQP 244
6**************************************************************************997 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 8.6E-33 | 162 | 229 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 31.419 | 165 | 242 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 9.03E-37 | 166 | 246 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 4.6E-29 | 168 | 241 | IPR004333 | Transcription factor, SBP-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005886 | Cellular Component | plasma membrane | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1050 aa Download sequence Send to blast |
MEDLGTAQVV SPTVIHQSMV GRFHDSYNPT SKKRGPPFHS SNSVHNKSPS DNWNPKSWNW 60 DSARFVAKPV QCDNGFDVGG GAPQIQIQTS LPARKEVLNG APIPRKPDRT GGDDENLRLK 120 LGGENGGGVN NNNTNNTNNN NNNTVEMQLR PSKRVRSGSP GGANYPVCQV DNCVKDLSAA 180 KDYHRRHKVC EFHSKAGNAL VGKQMQRFCQ QCSRFHPLSE FDEGKRSCRR RLAGHNRRRR 240 KTQPEDTVNA PQSLVPCARD NTVNDSDIVN LLAVLSRAQG NTEERSGKIP AIPDKDQLIQ 300 ILSKIHSLPA QTNMPSKPNG TVLNNVPSEN QNQINGKNNS STSTKNLLVA LSAHTSSQGS 360 DSEKSKSPCV DNNRDTIIVF PTVGGERSST SYHSPMEEVQ ETSPCVPLEL FSPSPEDYRS 420 MKLPSDGNFL SSGSSNPSVD RTPLSSPPVV YDLFPMRTMK DDCLSNNVGE IAYVKATMSN 480 GCSTSLQLFG SSKLATENGS IQSSPYRAGY ASSGSDHSPS SLNSDAQDRT GRIIFKLFDK 540 DPSHLPGSLQ TQIYSWLSNS PSEMESYIRP GCIVLSLYLS MPSFTWDQID ENLLNYVKSL 600 VKDVDIDFWG NGRFLVHTDR QRVSHKEGKI RLCKSWRTWN TPELIAVSPL AVVGGQETSL 660 LLRGRSLTAP GTMIHCTHAT GYNINEVPLS QDTPFDEVTL ACFKVNGTLG RCFIEVENNF 720 KGTSFPVIIA NNTICQELRL LEPEINGTAG VSDGIYREKA LGFLDELGWL FQRKQNSFLF 780 GIPDYRINRF KFLLIFSVEH DFCALVKTLL DILLELNLGR KGLLEKESLE LLSEIHLLNR 840 AVKRRCLSMV DLLVRYSVID SSEASGKFFF TPDMAGPGGI TPLHLAACTP SSDDMVDALT 900 SDPQKIGLQS WNTALDANGL SPYAYALMTN NHSYNALVAR KIADKENGQV SLSIENEIVQ 960 SQSEVDKRDK AISTFNQTQK SCSKCALAVR VHNSKKKFSG SKGLLQRPYI HSMLVVAAVC 1020 VCVCVFLRGH PYVGCVSPFA WENLGYGAI* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 4e-33 | 158 | 241 | 1 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 839 | 844 | RAVKRR |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' (By similarity). May play a role in plant development. {ECO:0000250, ECO:0000269|PubMed:15703061}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00664 | PBM | Transfer from Potri.002G002400 | Download |
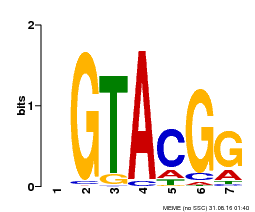
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Migut.C00339.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012832811.1 | 0.0 | PREDICTED: squamosa promoter-binding-like protein 14 | ||||
| Refseq | XP_012832812.1 | 0.0 | PREDICTED: squamosa promoter-binding-like protein 14 | ||||
| Swissprot | Q8RY95 | 0.0 | SPL14_ARATH; Squamosa promoter-binding-like protein 14 | ||||
| TrEMBL | A0A022RMB0 | 0.0 | A0A022RMB0_ERYGU; Uncharacterized protein | ||||
| STRING | Migut.C00339.1.p | 0.0 | (Erythranthe guttata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA12887 | 12 | 19 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G20980.1 | 0.0 | squamosa promoter binding protein-like 14 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Migut.C00339.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



