 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Migut.B01702.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Phrymaceae; Erythranthe
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 805aa MW: 87294.9 Da PI: 6.2881 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 63.6 | 2.8e-20 | 158 | 213 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t+ q++eLe+lF+++++p++++r eL+++l+L++rqVk+WFqNrR+++k
Migut.B01702.1.p 158 KKRYHRHTPLQIQELESLFKECPHPDEKQRLELSRRLSLETRQVKFWFQNRRTQMK 213
688999***********************************************999 PP
| |||||||
| 2 | START | 21.6 | 4.4e-08 | 356 | 434 | 1 | 69 |
HHHHHHHHHHHHHHHHC-TT-EEEE....EXCCTTEEEEEEESSS.......SCEEEEEEEECCSCHHHHHHHHHCCCG CS
START 1 elaeeaaqelvkkalaeepgWvkss....esengdevlqkfeeskv......dsgealrasgvvdmvlallveellddk 69
ela++a++elvk+a+ +ep+W + e++n+de ++ ++s + +ea+r++g+v+ + lve+l+d++
Migut.B01702.1.p 356 ELALAAMDELVKLAQTDEPLWIRGFeggkEILNRDEFSRTCAPSIIgmkpngFVTEASRETGMVIINGLALVETLMDSM 434
5899*******************9999999***********999989******************************98 PP
| |||||||
| 3 | START | 124.3 | 1.5e-39 | 434 | 543 | 94 | 206 |
EEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--.-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SSXXHHHHHHHH CS
START 94 mvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppesssvvRaellpSgiliepksnghskvtwvehvdlkgrlphwllrslv 185
m+a+lq+lsplvp R++ f+R+++q+ +g+w++vd S+++ pp + +++lpSg+++++++ng+skvtwveh++++++++h+l+r+l+
Migut.B01702.1.p 434 MNAALQVLSPLVPvREVNFLRFCKQHAEGVWAVVDLSIENLGGPP----FPTCRRLPSGCVVQDMPNGYSKVTWVEHIEYDESVIHQLYRPLI 522
789*****************************************9....666778************************************** PP
HHHHHHHHHHHHHHTXXXXXX CS
START 186 ksglaegaktwvatlqrqcek 206
+ g+ +ga +wvatlqrqce+
Migut.B01702.1.p 523 SAGMGFGAHRWVATLQRQCEC 543
*******************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 2.78E-20 | 145 | 215 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 6.2E-21 | 153 | 223 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.329 | 155 | 215 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 2.2E-18 | 156 | 219 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.52E-18 | 157 | 215 | No hit | No description |
| Pfam | PF00046 | 7.6E-18 | 158 | 213 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 190 | 213 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 28.475 | 347 | 546 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 3.25E-25 | 349 | 543 | No hit | No description |
| CDD | cd08875 | 3.26E-89 | 351 | 542 | No hit | No description |
| SMART | SM00234 | 1.1E-21 | 356 | 543 | IPR002913 | START domain |
| Pfam | PF01852 | 5.5E-34 | 434 | 543 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 1.05E-23 | 571 | 796 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 805 aa Download sequence Send to blast |
MNFGDFLDSN SCGGGGGGGG GGGGSRIVAD ITYTNNNNHN NNNASIVTTI PAGSNNIIMP 60 TGAISHPRLL PHSSLSTTTT TTKPMFLSPG LSLALQTNME GQGEIGRMVE KSEIMSNNNN 120 VGGRRSREEE HDSRSGSDNF DGGASADELD ASDKPPRKKR YHRHTPLQIQ ELESLFKECP 180 HPDEKQRLEL SRRLSLETRQ VKFWFQNRRT QMKTQLERHE NSILRQDNDK LRAENMSIRE 240 AMRNPMCTNC GGPALIGEIS LEEQHLRIEN ARLKDELERV CSLAGKFMGR PVSSMGHPMP 300 NSSLELGVGG INVISSDFGI SSPIPMLPPH KSITMSNVAT NNNNNDRSFE RSMYLELALA 360 AMDELVKLAQ TDEPLWIRGF EGGKEILNRD EFSRTCAPSI IGMKPNGFVT EASRETGMVI 420 INGLALVETL MDSMNAALQV LSPLVPVREV NFLRFCKQHA EGVWAVVDLS IENLGGPPFP 480 TCRRLPSGCV VQDMPNGYSK VTWVEHIEYD ESVIHQLYRP LISAGMGFGA HRWVATLQRQ 540 CECLAILMSS TSPATDHHTA ITAGGRRSML KLAQRMTNNF CAGVCASSVH KWNKLRTENV 600 DDDVQVMTRK SVDDPGEPPG IVLSAATSVW LPVTPQRVFD FLRDERLRSE WDILSNGGPM 660 QEMAHIAKGQ DHGNCVSLLR AGAVNSSQSS MLILQETCID AAGALVVYAP VDIPAMHLVM 720 NGGDSAYVAL LPSGFSIMPD GTGGGGPVSG DGSGGNGGSL LTVGFQILVN SLPTAKLTVE 780 SVETVNNLIS CTVQKIRAAL HCDN* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
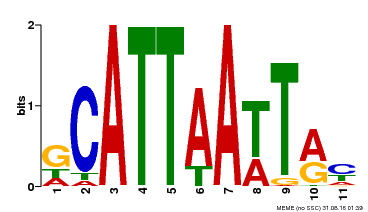
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Migut.B01702.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012843673.1 | 0.0 | PREDICTED: LOW QUALITY PROTEIN: homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | A0A2G9HWV6 | 0.0 | A0A2G9HWV6_9LAMI; Transcription factor DLX | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA1626 | 24 | 65 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00730.1 | 0.0 | HD-ZIP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Migut.B01702.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



