 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Manes.13G013200.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Euphorbiaceae; Crotonoideae; Manihoteae; Manihot
|
||||||||
| Family | ARR-B | ||||||||
| Protein Properties | Length: 584aa MW: 65471.9 Da PI: 6.1035 | ||||||||
| Description | ARR-B family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 77 | 2.4e-24 | 200 | 253 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
k+r++W+ +LH++Fv+av+q+ G +k Pk+il+lm+v+ Lt+e+v+SHLQkYRl
Manes.13G013200.1.p 200 KARVVWSVDLHQKFVKAVNQI-GFDKVGPKKILDLMNVPWLTRENVASHLQKYRL 253
68*******************.9*******************************8 PP
| |||||||
| 2 | Response_reg | 75.5 | 2e-25 | 20 | 128 | 1 | 109 |
EEEESSSHHHHHHHHHHHHHTTCEEEEEESSHHHHHHHHHHHH..ESEEEEESSCTTSEHHHHHHHHHHHTTTSEEEEEESTTTHHHHHH CS
Response_reg 1 vlivdDeplvrellrqalekegyeevaeaddgeealellkekd..pDlillDiempgmdGlellkeireeepklpiivvtahgeeedale 88
vl+vdD+ + +++l+++l+k y ev+++ + +al+ll+e++ +D+++ D++mp+mdG++ll++ e +lp+i+++ ge + + +
Manes.13G013200.1.p 20 VLVVDDDLTWLKILEKMLKKCSY-EVTTCGLAIDALHLLRERKdrYDIVISDVNMPDMDGFKLLEHVGLEM-DLPVIMMSVDGETSQVMK 107
89*********************.***************888778**********************6644.8***************** PP
HHHTTESEEEESS--HHHHHH CS
Response_reg 89 alkaGakdflsKpfdpeelvk 109
++ Ga d+l Kp+ ++el++
Manes.13G013200.1.p 108 GVQHGACDYLLKPIRMKELRN 128
******************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PIRSF | PIRSF036392 | 7.6E-147 | 1 | 547 | IPR017053 | Response regulator B-type, plant |
| SuperFamily | SSF52172 | 5.27E-35 | 17 | 147 | IPR011006 | CheY-like superfamily |
| Gene3D | G3DSA:3.40.50.2300 | 1.8E-43 | 17 | 141 | No hit | No description |
| SMART | SM00448 | 7.2E-31 | 18 | 130 | IPR001789 | Signal transduction response regulator, receiver domain |
| PROSITE profile | PS50110 | 42.771 | 19 | 134 | IPR001789 | Signal transduction response regulator, receiver domain |
| Pfam | PF00072 | 9.4E-23 | 20 | 129 | IPR001789 | Signal transduction response regulator, receiver domain |
| CDD | cd00156 | 6.52E-29 | 21 | 134 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 7.8E-28 | 198 | 258 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 4.3E-17 | 198 | 257 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.7E-21 | 200 | 253 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.8E-7 | 202 | 252 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000160 | Biological Process | phosphorelay signal transduction system | ||||
| GO:0009735 | Biological Process | response to cytokinin | ||||
| GO:0010082 | Biological Process | regulation of root meristem growth | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 584 aa Download sequence Send to blast |
MMDNGFSSPR HDPFPAGLRV LVVDDDLTWL KILEKMLKKC SYEVTTCGLA IDALHLLRER 60 KDRYDIVISD VNMPDMDGFK LLEHVGLEMD LPVIMMSVDG ETSQVMKGVQ HGACDYLLKP 120 IRMKELRNIW QHVLRKKIHE VRDIEILEGM ESIQMTRNGL DQSSDGHLLC GEDLTSVKKR 180 KEAENKHDDK DPGDSSSTKK ARVVWSVDLH QKFVKAVNQI GFDKVGPKKI LDLMNVPWLT 240 RENVASHLQK YRLYLSRLQK ENDTKTSVGG GKHSDSPSRD SAGSFGIQNS INIQRNDISN 300 GSYGFSGNSL IVRNVEPRSQ ENERKGIVSK TAVEPKRALA VEVPDPCKPK SSEMEFGHSF 360 TSPASEVNFA EFGSNFPTKF SWCAIPQSQL KQEQNPLHLD AGFSQRTRPG KQQHIQVDYP 420 QPSPPIISGS SVTERNMDGS VKIKPIYDEC RNNGSQVSSA RSTMDSYQVQ TKTYEVNHQA 480 YEPISTNTSS LKNQAAFNLS SISDLESAQK SINWAMSPLA TLDDDFQVRW VQGDCYAMNL 540 GLQNIEFPEY FDGGLLADVP THLYENLYDS TECSVIEQGL FIA* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 2e-17 | 196 | 258 | 1 | 63 | ARR10-B |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00010 | PBM | Transfer from AT1G67710 | Download |
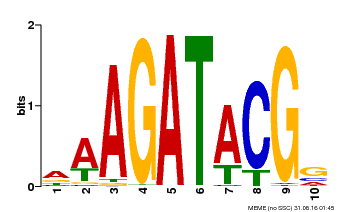
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Manes.13G013200.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021633533.1 | 0.0 | two-component response regulator ARR11-like | ||||
| TrEMBL | A0A2C9UNZ6 | 0.0 | A0A2C9UNZ6_MANES; Two-component response regulator | ||||
| STRING | cassava4.1_004317m | 0.0 | (Manihot esculenta) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF3999 | 33 | 60 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G67710.1 | 1e-131 | response regulator 11 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Manes.13G013200.1.p |



