 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Manes.09G109300.2.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Euphorbiaceae; Crotonoideae; Manihoteae; Manihot
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 368aa MW: 41367.8 Da PI: 8.7426 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 101.1 | 7.3e-32 | 63 | 116 | 2 | 55 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
prlrWtpeLH +Fv+ave+LGG+e+AtPk +l+lm+v+gL+++hvkSHLQ+YR+
Manes.09G109300.2.p 63 PRLRWTPELHLCFVKAVERLGGQERATPKLVLQLMNVNGLSIAHVKSHLQMYRS 116
8****************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 1.7E-30 | 59 | 117 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 14.255 | 59 | 119 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 6.27E-15 | 61 | 117 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 3.8E-23 | 63 | 118 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 7.2E-10 | 64 | 115 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 368 aa Download sequence Send to blast |
MEGSDKTGCS KTRACRQNQD ESESGENDED GNKVKNGGSS SNSTVEESEK KSSVRPYVRS 60 KMPRLRWTPE LHLCFVKAVE RLGGQERATP KLVLQLMNVN GLSIAHVKSH LQMYRSKKID 120 DPSQAMADHR HLVESGDRNI YNLSQLPMLQ GYNQRHGSSY RYGDASWNSR ESFVYNPHMG 180 RCLVDETRQG FYGTVAERIF GGNNTSNWTN CKLQTGASSF RSQFNWKAEE QLKAEQRRPS 240 QNPKFWQTQP SSSFTQLNPA AQVLNRSNIN IFSDMKSETN LQELKSLKRK ATDCNLDLDL 300 SLKLTPASDG SRRSLEESEV DSELSLSLYS PSSSKLSRLK GLGEDDYNNN SNNKEDAKRA 360 STLDLTI* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4r_A | 1e-18 | 64 | 118 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 1e-18 | 64 | 118 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 1e-18 | 64 | 118 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 1e-18 | 64 | 118 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
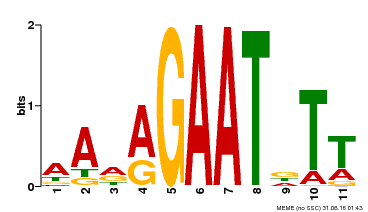
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00297 | DAP | Transfer from AT2G38300 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Manes.09G109300.2.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021622642.1 | 0.0 | putative Myb family transcription factor At1g14600 isoform X2 | ||||
| TrEMBL | A0A251K5N0 | 0.0 | A0A251K5N0_MANES; Uncharacterized protein | ||||
| STRING | cassava4.1_026842m | 0.0 | (Manihot esculenta) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G38300.1 | 9e-60 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Manes.09G109300.2.p |



